|
My internship project is the estimation from genetic data of the long term effective population size (Ne) of human populations from the Austro-asiatic linguistic group located in South-East Asia. This value, sometimes described as the number of “independent breeding individuals” represents the level of genetic drift that led to the current genetic diversity. Information from population mating system, life and demographic history can be inferred from Ne.
I will use two different Linkage Disequilibrium (LD)-based methods to get two Ne estimators including one from which I can infer its variations over generations as well as the time since divergence between two populations, allowing me to confront my results with regional settlement scenarios. This summary is the description of my project as it is planned on the 21st of march 2017 and can therefore be subject to change especially concerning the second method which I am currently evaluating in terms of time and computational cost (in one word : feasibility !). Picture's source : http://www.earth.com/news/human-population-exploded-animation/
4 Comments
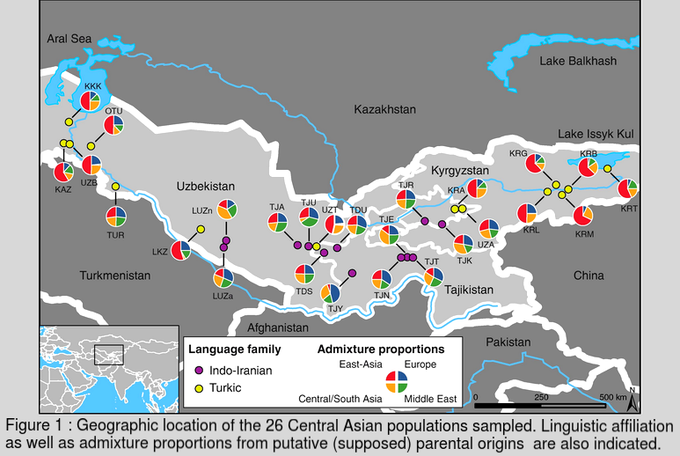
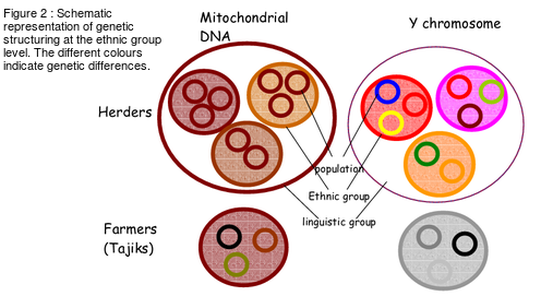
This first article of my internship blog is an introduction to the research field of population genetics and especially to the approaches developed by the evolutionary anthropology research team located in the “Museum of Mankind” (Musée de l’homme), Paris (1). Genetic variations between individuals, populations, linguistic groups are studied by population geneticists all around the world. Researchers can deduce from data of the distribution of alleles in a studied populations, many aspects of these populations demography or adaptation (2). I will present you today some demographic examples : from the relative distances, levels of admixture to the histories or even cultures of human populations ! Genetic diversity is the result of evolutionary forces As I mentioned it above, the majority of our knowledge on human genomes evolution rely on the study of genetic diversity between individuals also called polymorphism. So it seems logical to remind you the four major evolutionary forces that lead to the modification of allele frequencies in a population future generations (see Box 1)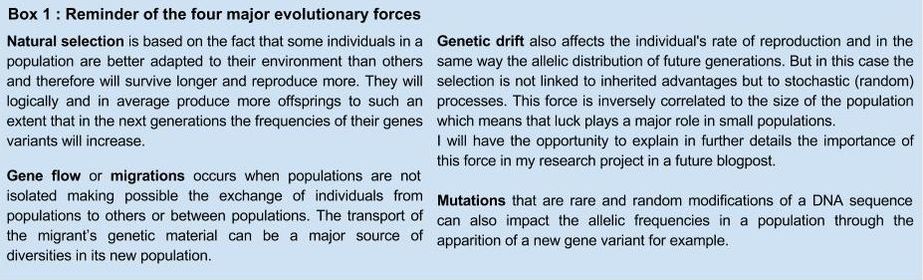  The understanding of the relative strengths of these process (as well as other biological process such as recombination that have an impact on the linkage probability between two different genes on the same chromosome) allows the scientists to draw conclusions on present or past changes from genetic data. Genotyping give access to the personal information stored in our DNA DNA is sampled from related or/and unrelated individuals (both are useful in my work) inside the studied population. Subjects must give their informed consent to the researchers and DNA sample is completed by a personal survey on their familial background, ethnicity etc The whole genome is rarely genotyped and the research team agrees on what kind of neutral or non-neutral genetic markers should be used in further researches (3). I personally use some markers called SNPs (Single Nucleotide Polymorphisms) which are variations at a particular and identified position of a single nucleotide. We can therefore know the nucleotide (between A, T, C or G) that each of the sampled individual carries at dozens of thousands positions distributed along the chromosomes. Study of genetic diversity in Central Asian populations at different scales The example, I am going to quickly describe is part of the laboratory research work on the question of Central Asian populations genetic diversity. Summary of the complex ethnographic situation in Central Asia There are two linguistic groups associated with different livelihoods, the Indo-iranian are sedentary agriculturalists whereas the Turkic speakers are mainly nomadic herders (shepherd). Between all kinds of analyses that have been realized on the sampled populations, I present you here the results of the admixture analysis that as well as the sex-specific marker diversity one which give us information respectively on population ancestry or settlement and culture. Data analyses : admixture proportions from parental populations (4) It is possible to obtain for each population the maximum likelihood estimation of the proportion of the genome from four supposed parental (well known) populations (located in East Asia, Europe, Middle East and Central/South Asia) (Figure 1, Martínez-Cruz, Begoña et al, 2010). This analysis permit to obtain a genetic landscape between linguistic and ethnic groups in this region. As you can easily observe it on the map, the research team was able to conclude that Indo-iranians speakers were closer to Western Eurasians contrary to Turkic who are closer to East-Asians. Data analyses : sex-specific markers diversity (5) As you may know, mitochondrial DNA is transmitted from mother to children whereas Y chromosome is transmitted from father to son. The study of genetic markers on these two DNA sequences permit to understand the specific demographic histories of men and women in this region. Indeed the team calculated the differentiation rate (Fst value) between populations of these uniparental markers and get very different results. They observed that Y chromosome DNA was much more diverse than mitochondrial DNA between herders populations (villages). This observation can be explained as the results of a cultural rule called exogamy and characterized by the marriage of women outside their birth-village. Different villages have then been overtime genetically more separated in term of male genomes (men in one village are relatives) than female one (figure 2, Heyer, Evelyne et al, 2009). Results of the analyzes of intra/inter groups diversity and conclusions : Admixture analysis seems to show that the two linguistic groups are the structure of the main differentiation between populations. However the results of sex-specific markers analysis which targets more precise variations highlight the existence of different levels of genetic diversity in the genomes of individuals from the same linguistic group. This results can lead in one hand (admixture) to a new regional settlement hypothesis and in the other hand (sex-specific markers) to the questioning of the way we perceive culture as a result and not as the origin of differentiated genetic groups. Take-home message ! To be fully honest, this example, that I personally find very interesting is not exactly related to my internship project. Indeed I am working neither on the same populations nor on the same kind of sex-specific markers analysis. I will obviously write about what I concretely do in my next few posts, but now I would like to explain why I chose to describe this study in particular. This population genetics study has began years ago and is now at a very advanced stage. So it can be a good overview of the possibilities of genetic data analysis in the understanding of the close link between culture and genome as well as in historic scenarios reconstruction. By reading this first blog you should be much more able to understand the issues of my work that I am going to present you from the most theoretical to the most practical steps in this blog ! Bibliography
(1) http://www.ecoanthropologie.cnrs.fr/ (2) Balaresque, P. L., S. J. Ballereau, and M. A. Jobling. "Challenges In Human Genetic Diversity: Demographic History And Adaptation". Human Molecular Genetics 16.R2 (2007): R134-R139. Web. (3) Vignal, Alain et al. "A Review On SNP And Other Types Of Molecular Markers And Their Use In Animal Genetics". Genetics Selection Evolution 34.3 (2002): 275-305. Web. (4) Martínez-Cruz, Begoña et al. "In The Heartland Of Eurasia: The Multilocus Genetic Landscape Of Central Asian Populations". European Journal of Human Genetics 19.2 (2010): 216-223. Web. (5) Heyer, Evelyne et al. "Genetic Diversity And The Emergence Of Ethnic Groups In Central Asia". BMC Genetics 10.1 (2009): 49. Web. |