|
Everybody hello! I must say that I’ve been quite busy those 2 last months organizing night follow experiments on dinospores’ fate. Working in a laboratory implies protocol reviews and team management, which has been a big and great practice during past weeks! So here we are, after calibration and good maintaining of targeted cultures I finally started. Table 1. gather all cross-infection I worked on (for further understanding, refer to pasts article: An idea for Amoebophrya’s dinospores maturation time?). Experiment 1: Optimisation of incubatin time First part of the protocol was to obtain details on how long both parasite strains need to be in contact to initialize an infection in ST147 (Scrippsiella trochoidea). I designed a protocol based on previous results (refer to: An idea for Amoebophrya’s dinospores maturation time?) that have shown that maturation time for dinospores of strain A25(ST) is around 18h and 6h for A120(ST) in host ST147. '>  Table 1. Incubation time for strains A25(ST) and A120(ST) to initialize host infection The minimal time required for infection was checked using incubations from 15 minutes to 3 hours. Step 1 consisted on disnospore production: filter cultures onto 5 µm polycarbonate filters and inoculated on a new fresh host ratio 1:10. Step 2 consisted of collecting dinospores after a complete infection cycle: filter onto 5 µm filters with autoclaved devices and wait maturation time according to previous results (18h for A25(ST) and 6h for A120(ST)). Step 3: inoculation of dinospores on a new fresh host culture and collect samples each 15 minutes for 1h and each hour for 3h. We repeated the test with the A25(ST)*ST147 to increase this maturation period (24 hours). Experiment 2: Fate and infectivity of dinospores Next part consisted on collected samples to follow dinospores fate: maturation and infectivity. This protocol was also designed regarding previous FISH results (blogpost from 11/11) and results from a). Step 1 consisted on dinospore production similar to a) but ratio 1:3 to boost infection. Step 2 is proceeded 24h incubation later for reduction of the bacterial contamination. Infected host cells are collected using 10 µm nylon filters, then resuspended into 30 mL of fresh medium (recently filtrated again onto 0.2 µm) twice. Second time, we filter 200mL media to rinse cells. Cells are resuspended into 560 mL of fresh medium. This culture was then split into 6 different flasks (90 mL each). In three of them, antibiotics (PNS: Penicillin-Streptomycin-Neomycin), were added (3% final concentration). Step 3 consists of dinospore production and monitoring. After 24 hours, the culture was checked every 6h for dinospore production. Freshly produced dinospores were gathered by 5 µm PC filtration. Samples were collected for flow cytometry (signatures and ploidy levels) analyses and DAPI microscopy each 2 hours during 22h for A25(ST) and each hour for 8h for A120(ST). At same times, a sample was used to reinfect fresh host cultures at a final ratio of 1:10 (parasite: host) for A25(ST) and A120(ST). Incubation times differ over strains and were determined based on Experiment 1. Cultures were then fixed with PFA (1% final concentration) for FISH analyses.
0 Comments
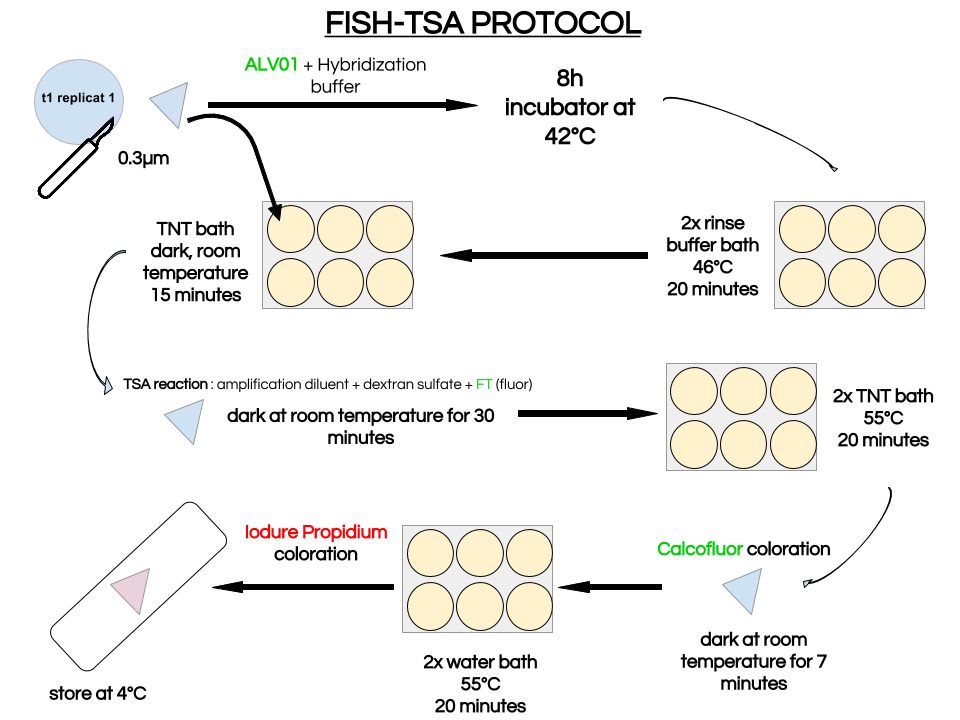
Dear all, Since the beginning of my internship, our subject has not stopped to evolve. Now that I’m halfway through, our interest has finally found a stand : what happens when Amoebophrya ceratii complex species dinospores coming from a previous host infection are released in the column water until they infect a new host ? This general question implies 2 related points. Does dinospores become mature ? Do they acquire their virulence ? Some of those hypothesis were already investigated in 2013 by some members of the team butshould be done again with a refine experimental protocol. Those previous results on maturation time have shown different dinospore populations over time after their release by flow cytometry and set us on the right path for further experiments. In fact, for some other parasitic dinoflagellate species such as Euduboscquella crenulata, 3 different dinospore morphotypes are known and probably linked with its sexual reproduction whilst for another one, Ichtyodinium chaberlarli that also produces different dinospore morphotypes. This phenomenon was linked to different division generations. So my first task here was to focus on dinospores maturation by analysing FISH filters. FISH (Fluorescence in situ hybridization) here is a method that allow the labeling of one specific organism thanks to oligonucleotide probes attached by a fluorochrome. 5 different Amoebophrya strains have been put on 0.3µm filters at different times and I had to stain them with a probe ALV01, specific for the parasite and 2 other dyes: Calcofluor and Iodure Propidium that stained the host theca and nuclei of both partners, respectively. We could then analyse filters and separate hosts and parasite for counting different infection stages and total infection prevalences thanks to epifluorescence microscopy. The different stages are : 0 : no infection 1 : initial infection in the cytoplasm 2 : at least one parasite in the nucleus, part of the nucleus remains 3 : the host nucleus is totally digested but part of the host cytoplasm remains 4 : the whole host is digested (multinucleated vermiform with intact host theca) Amoebophrya strains here studied have particularities :
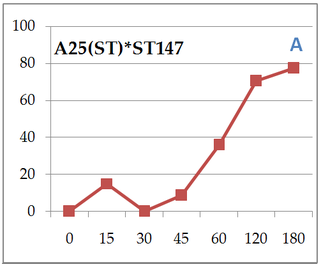
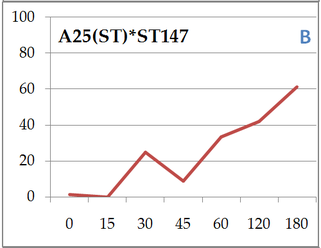
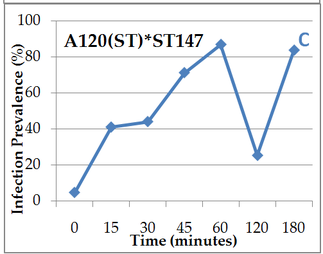
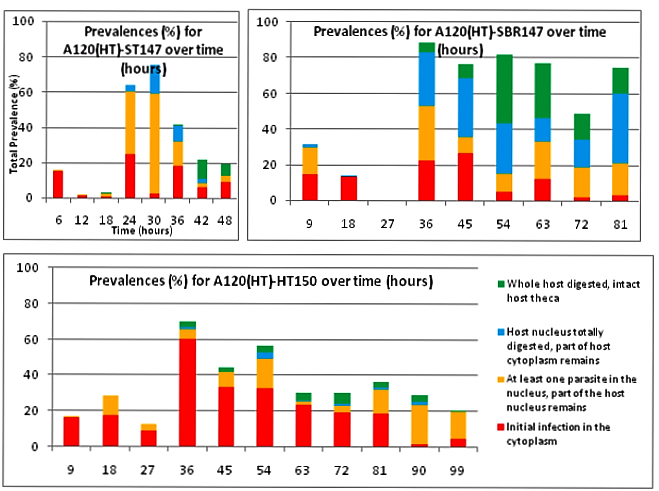
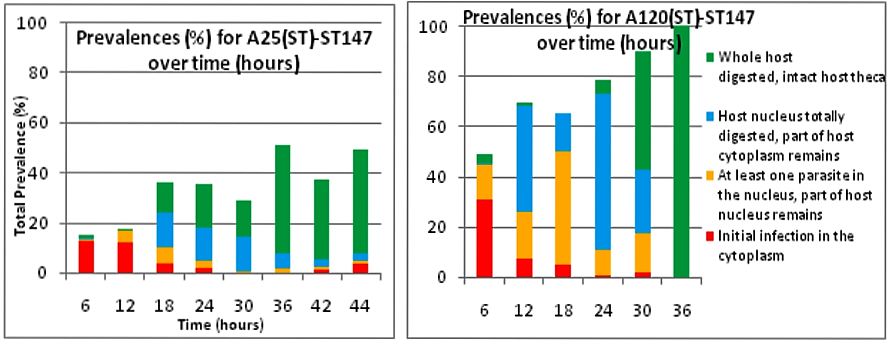
Here we can compare 2 different strains A120 and A25. Both parasitic strains have been purified 5 times in 2010 (clonal strain) before sequencing their complete genome and get their transcriptomes over a complete infection cycle. The strain A120 was able to infect different host species, and for security reason (to not lost it) was maintained into two different host species (Heterocapsa triquetra= HT and Scrippsiella trochoidea=ST). 5.5 years after, the project was to obtain the transcriptome of this strain infecting different host species. The strain maintained in ST (A120(ST)) was not able to infect HT anymore. The strain maintained in ST (A120(ST)) was able to infect S. trochoidea, S. donghaeinis and Heterocapsa triquetra. This strain was used to monitor the infection dynamic and collect transcriptome during the complete life cycle. I was in charge of acquiring data for dynamic of infection by FISH, and to compare this dynamic and phenotypes over host strains. Comparison of one strain in different hosts, A120(HT): Infection dynamic is different depending on the host as for the 3 described hosts, dinospore maturation takes different times. It seems that in none of this 3 cases the parasite finalize its infection as few stage 4 (whole host digested with theca) is observed at the end of infection cycle and as general tendencies of each graphs correspond to a decrease. This decrease point the fact that although infection is going on, uninfected hosts keep dividing and win the battle against the parasite. Comparison of one strain from different hosts in the same host Scrippsiella trochoidea ST147: Same strain A120 cultivated in 2 different hosts HT and ST duri and put under same condition to infect a same host ST147 don’t have the same infection dynamic. Their phenotypes for infection aren’t similar, as dinospores did not seem to be able to infect a novel host for A120(HT)-ST147 before 24h whereas it is 6h for A120(ST)-ST147. This is supported by the fact that A120(HT)-ST147 does not succeed in finishing infection process as the general decreasing tendency and the soft prevalence for stage 4 (whole host digested with theca remaining) at time 42 and 48 suppose the host continues to divide. On the contrary, 100% total infection prevalence is reached at time 36 for strain A120(ST)-ST147 with dominant stage for at time 30 and 36. In this case the parasite succeeds its infection process. We can conclude that phenotype here is not strain dependent but as before, host dependent as same strain in the same host but coming from different previous host do not have the same behavior. Comparison of 2 strains A120 and A25 from same host Scrippsiella trochoidea in the same host Scrippsiella trochoidea ST147: For different strains from same host A120(ST) and A25(ST) in a same host ST147, phenotypes aren’t the same considering dinospore maturation time. But it is interesting to notice that on the contrary as for A120(HT)-ST147 and A120(ST)-ST147, A120(ST)-ST147 and A25(ST)-ST147 succeed in finalizing their infection cycle. Once again we proved that phenotypes aren’t the same for different strains even if here they have been cultivated the same exact way.
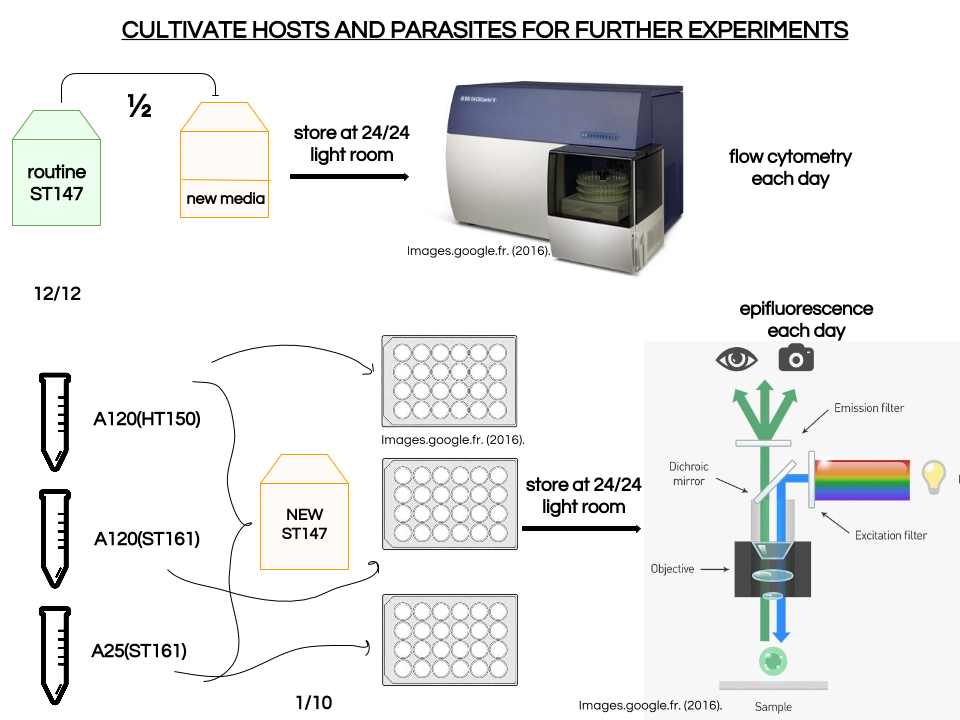
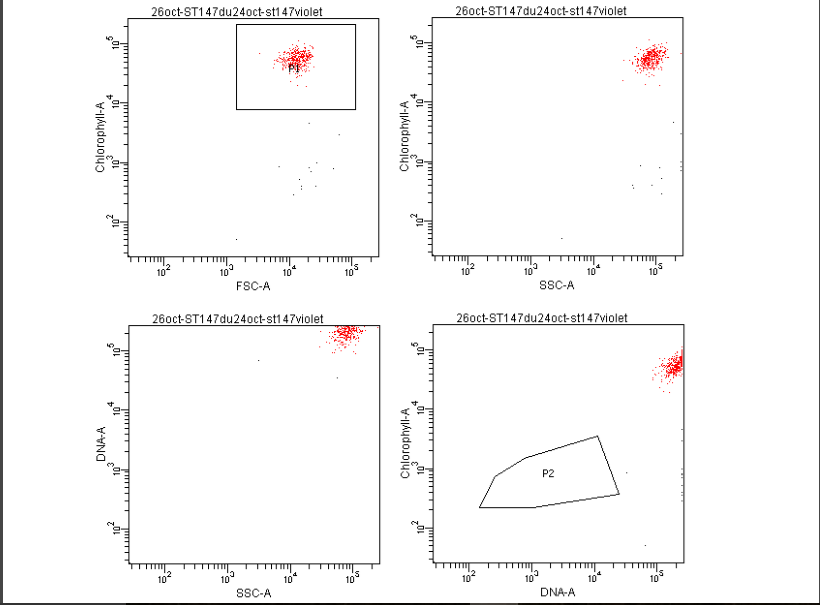
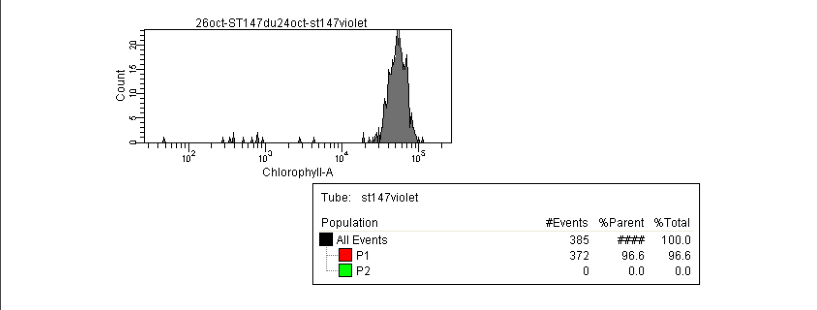
We also notice that A25(ST)-ST147 seems to be stuck at stage 4 (whole host digested with remaining theca) from time 18 to time 44. A possible explanation for this observation is the host encystment that prevent the release of the parasite which lead to this delay in the infection process. Here we have demonstrated that there is dinospore maturation over times depended on hosts targeted and not on strain concerned. We now have an idea of how long takes this maturation for further experiment on A120(HT)-ST147 dinospore maturation : is there different population of dinospores through this maturation corresponding to different stages (fusion, division) ? Till’ next time, this give pause for thought, Bye ! Hi y’all, (sorry there was suppose to be a post before this one but I’m waiting for confidential validation...) This blogpost is to catch your attention on how it is important to well prepare an experiment. Through those 2 past weeks, I have experienced bacterial contamination in my host culture ST147. For the big day experiment, I needed to change environmental conditions than as routine. My host ST147 was usually cultivated in light 12/12 meaning 12 hours light and 12 hours night per day and pricking out each monday and friday. For further experiment on dinospores maturation, we needed to change light to 24/24 which means 24 hours of light per day and as for routine, pricking out each monday and friday. This changes allow the host to grow faster in the media, the aim being to have 8000ȼ (cells/mL) host each monday and friday for pricking out our 3 strains of interest : A120(HT)-ST147, A120(ST)-ST147 and A25(ST)-ST147 described in previous/following posts. So for the host we took the usual flask containing ST147 for routine and prick out on monday with new media ratio ½. Then we store the culture in a room 24/24. For the strains, we took routine A25 in ST161, A120 in ST161 and A120 in HT150 and put them in our new ST147 host culture ratio 1/10. Both host and parasites are prick out on monday and friday. Thanks to epifluorescence microscopy, we can follow the life cycle of parasites each day to be sure when dinospores are released after host infection, how long it takes and so when we have to prick out to adjust ratio etc... Thanks to flow cytometry, we can estimate the number of host cells per mL each day to then piece together the growth curve and predict when there will be 8000ȼ and adjust ratio for monday and friday as there is not the same time between pricking out. Flow cytometry allows us to estimate number of cells per mL. In a tube, we put 500µL of our sample that we fix with 5µL glutaraldehyde. This permit to fix our cells in the stage they are in the moment we want to analyse them (prevent cells from dying as tube will be placed on cytometer with no hood although we sample under biological hood). Then the cytometer will vacuum the sample in a tiny little conduit for a fixed time (60s in our case) at a medium rate. I had to measure and determine the flow rate before analysing my samples because it can change at time to another. For that, a media sample is analysed for 10 minutes and weighed before and after analysis : flow rate = (w1-w2)(g)/time(min) Then samples are analysed for 60s at medium rate giving total events recorded with the cytometer. To have number of cells : nbcells = (totalevents * dilution)/(flowrate * time) → here dilution is 1 because we did not diluted our sample and time is 1 because it is in minutes. Different graphs can be realized thanks to different specific parameters such as SSC-A (volume), DNA-A (host nucleus) FSC-A ( size) and also Chlorophyll-A. Size vs Chlorophyll-A and Volume vs Chlorophyll-A can be realized without markers and Volume vs DNA-A such as DNA-A vs Chlorophyll are realized with markers such as Sybr Green. The graph Chlorophyll-A vs Count is really useful because the more there are chlorophylls, the more cells there are. So I started to follow my cultures each day and one week later I realised that number of cells was stuck at ≈1000ȼ. Regarding the flask, we could observe a milky liquid that seems correspond to bacterial contamination. Normally, when host develop the culture is blurred but not milky. Another flow cytometry technique allowed us to confirm this contamination. As we use natural Penzé media (water from Britain, France) for host cultures, it is quite normal to have some bacteria but not that much. To have the ratio between host and bacteria in a 500µL sample we fix cells as usual with 5µL glutaraldéhyde and with 1/5000 Sybr Green (marker for bacteria). Instead of defining only one population, we determine 2 windows on parameters for 2 populations targeted. We would then have 2 total events recorded for 2 populations. As we can see on graphs, bacteria was saturated compared to host → bacterial contamination was real ! We consequently have to start again the processus : we created a new flask with ST147 to acclimate at 24/24 culture room.
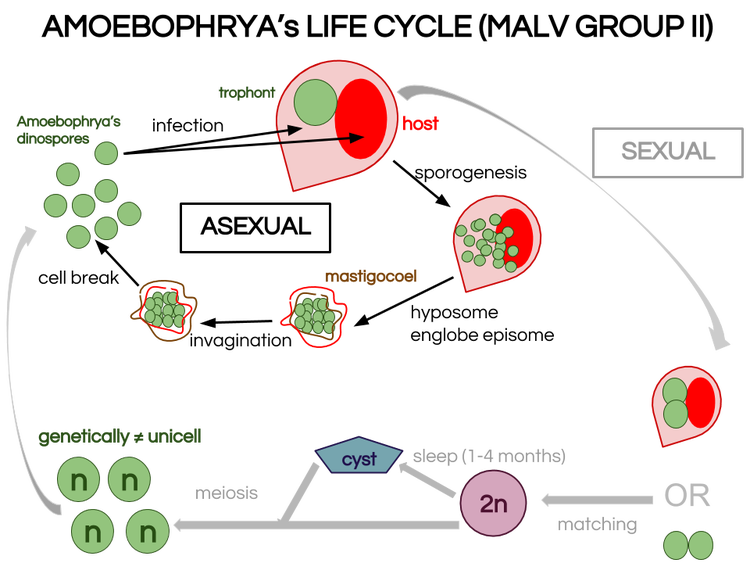
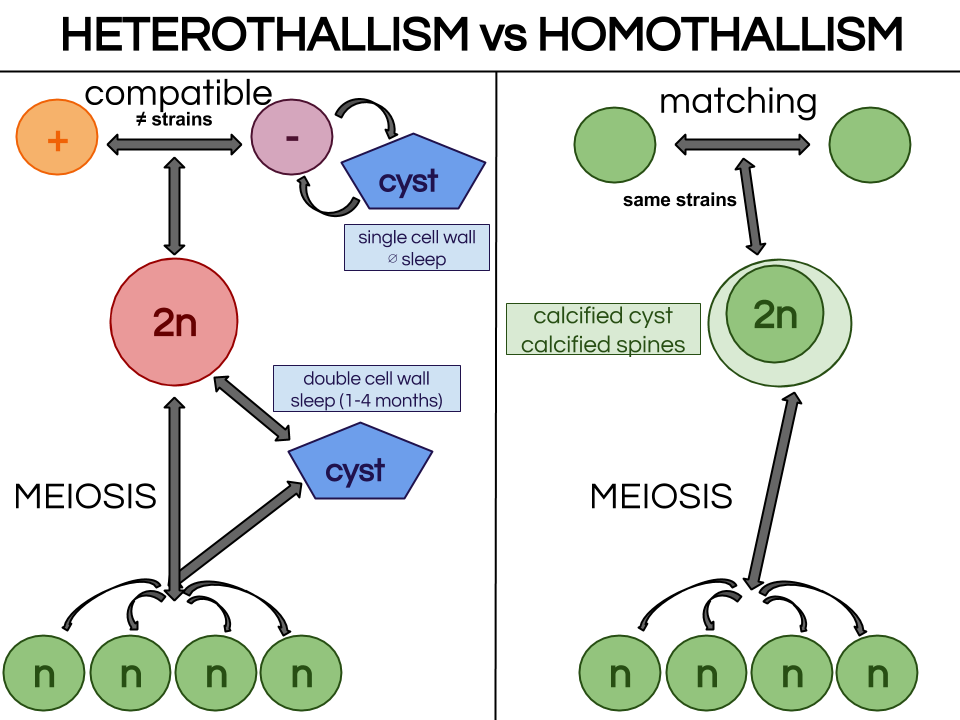
What I did not tell is as our host was not healthy because of that contamination, our parasites did not end up correctly their infection cycle so we had to start all over again for those bug too ! Now we wait until our hosts and parasite are really healthy for the big day experiment ! See y’all 😉 Demat ! (hello in Breton) Today I am going to demonstrate why Amoebophrya’s life cycle is so important in our study. Amoebophrya is from the dinoflagellates phylum and Syndiniales order, also known as MALV group II[1]. It contaminates toxic microalgae that are likely to create red-tide. When a host is contaminated, Amoebophrya prevent its reproduction and kill it to undergo their own life cycle, which can last from 2 to 4 days. Once finished, hundreds of new infectives parasites known as free-living dinospores are released. In this dispersive stage the parasite can survive only few days. Amoebophrya dinospores have typical morphology features such as their large bulbous episome and their narrow sharp hyposome. Their dense nucleus situated in the episome has peripherically condensed chromatin. Dinospore are biflagellates, each of them emerged from the episome base and are perpendicular to each other. The amphiesma is composed of a plasma membrane and a single layer of flat alveoli. Tricocysts, striated strip and 2 mitochondria appear to be important features involved in the entering process in the host cytoplasm/nucleus. The binding process implies the host cell surface, flagellar hairs, hyposome tip and a microtubular basket allowing dinospores to enter the cell. The previous general description actually corresponds to asexual reproduction, a vegetative life cycle which is nearly well-described. Let’s zoom in this complex asexual process. Endoparasitic vegetative life cycle : is Amoebophrya only asexual ?[2] It starts when a small dinospore invades the host cytoplasm and/or nucleus. In fact, dinospore are mostly intranuclear within thecated (theca is the hard mineralized envelope of several marine species) host species (Scrippsiella trochoidea) whereas they are intracytoplasmic within non-thecated host species (Heterocapsa triquetra). However, this general pattern is not endlessly observed. Consequently it become complicated to rely on this vegetative process as a sign of taxonomic feature for determination of species. Once in the cytoplasm or in the nucleus (which appear to have the same process as for the cytoplasm), the parasite becomes a small spheric mononuclear trophont. Through several flagellar and nuclear divisions, known as sporogenesis, the trophont becomes a multinucleate and multiflagellate beehive shape cell. In the meantime, the hyposome has expended and arched to englobe the episome forming a typical cavity, the mastigocoel[3]. Once mature, the beehive is transformed into a long vermiform by a complex process of invagination. The mastigocoel is inversed, which places flagella in the outer surface of the parasite and breaks host external membrane. The intern membrane becomes extern and vice-versa. The vermiform does not live for so long and breaks into hundreds infectious dinospores. During this cycle, an atypical feature for dinoflagellates is notable : chromosomes are not permanently condensed which differs from other dinoflagellates. Strains do not always infect the same specific host and their behaviour is really wide among the different clades. As one Amoebophrya strain may be both specific or generalist and as many exceptions appear through general descriptions, asexual life cycle cannot be considered to determine species within MALV Group II. Hence the wish to deepen our knowledge on sexual reproduction within this group. Life cycle dimorphism : what advantages sexual reproduction could give ? Sexual reproduction has not been observed frequently within dinoflagellates. Our goal is to determine if it occurs within Amoebophrya genus to separate this genus into species.[4] Moreover, life cycle dimorphism may convey several advantages to hosts or parasite depending on the cells released.[5] As Amoebophrya can be found in coastal planktonic ecosystems characterized by strong environmental fluctuations and rapid turnovers, production of diverse genetically different dinospores could allow survival of the parasite and prevent resistance disturbing marine microbial dynamic. In fact, if sexual reproduction occurs, genetic recombination occurs and meiose form 4 individual zygotes (n) different from the mother cell. As dinospore diversity may convey advantages for the parasite, it may also enforce host resistance. To survive, the host and the parasite need to evolve in resistance and virulence, respectively. Thus, the taking place of a huge fight within phytoplankton ecosystems. Our plan is to observe sexual reproduction or not within Amoebophrya strain A120 in Heterocapsa triquetra 150, SBR 147 (Scrippsiella donghaenis host) and ST 161 (Scrippsiella trochoidea host). If we observe matching, A120 is a homothallic strain, meaning that 2 same strain gametes can match. In this case, we will have to further observe how the stage 2n (diploïd) become n (haploïd) after gamete fusion in A120. If there is no matching within A120, the strain is heterothallic and we need to find which strain is compatible (not female and male matches) . For that, we will mix all strains from sub-clade 4 and look at any matches. If there are no matches there are 2 possibilities : mix A120 with all strains from another sub-clade or mix all strains of another subclade not regarding A120 anymore. A tantôt ! [1] Guillou, L. et al. "Widespread Occurrence And Genetic Diversity Of Marine Parasitoids Belonging To Syndiniales ( Alveolata )". Environmental Microbiology 10.12 (2008): 3349-3365. Web.
[2] Miller, John J., Charles F. Delwiche, and D. Wayne Coats. "Ultrastructure Of Amoebophrya Sp. And Its Changes During The Course Of Infection". Protist 163.5 (2012): 720-745. Web. [3] Cachon, Jean et Monique. "Ultrastructure Des Amoebophryidae (Péridiniens Duboscquodinida". Protistologica. 1st ed. 1969. Print. [4] Pagenkopp Lohan, Katrina M. et al. "Genotypic Variation In The Parasitic Dinoflagellate Hematodinium Perezi Along The Delmarva Peninsula, Virginia". Marine Biology 161.2 (2013): 261-273. Web. [5] Figueroa, Rosa I. et al. "The Hidden Sexuality Of Alexandrium Minutum: An Example Of Overlooked Sex In Dinoflagellates". PLOS ONE 10.11 (2015): e0142667. Web. Dear Prom', It has been one month since the beginning of my internship in the Station Biologique of Roscoff in the team DIPO, mainly working on interactions within phytoplankton. Our interest stands on Amoebophrya, a dinoflagellate parasite that infects other species from its phylum[1]. As this may be completely new to most of you, I'll start this first insight on the subject by definitions ! 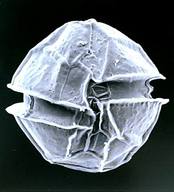 Figure1. Alexandrium minutium, another dinoflagellate specie (microscope x5700) Figure1. Alexandrium minutium, another dinoflagellate specie (microscope x5700) Symbiotic association As none of living species lives totally isolated from others, biological interactions are considered as the main force for evolutionary processes. Among these biological interactions, symbiotic associations are characterized by the sustainability of a specific relationship between two different species, kept from one generation to another. These two partners are called the host (the one that is hosting the second species) and the symbiont (the one that is living inside or on the host). The host can be used by the symbiont (parasitism) or, on the contrary, the symbiont can be used by the host (endosymbiosis). These effects are often reversible (an endosymbiont can suddenly act as a parasite and reciprocally), and all possible effects on the host (from lethal to beneficial) are observed in Nature. Parasitism Until the middle of XXème century, parasitism was only considered if there was a unilateral need for the parasite to infect a host in order to survive. Generally, the parasite is smaller than its host. In this larger definition, viruses are thus considered as parasites. The term “parasitoids” is used for parasites that kill their host to accomplish their life cycle. For unicellular, it is sometime difficult to distinguish parasitoids from predators (or grazers). Both kill their host/prey, both may have similar functions in Nature, by controlling their host/prey populations. For instance, Ameobophrya is a parasite that controls some microalgae species (Scrippsiella) that are likely to create red-tides (Figure2). However, there is a big difference between a predator/grazer and a parasite. A predator/grazer generally consumes several preys during its life and its division release two daughter cells. In contrast, a parasite feeds generally on a single host and produces more than two daughter cells during sporogenesis. This sporogenesis will ends by the production of > 2 spores, which are released outside the host and used to infect a novel host. Phytoplankton Our host and parasite are part of the primary food source known on Earth as phytoplankton. The phytoplankton is composed by unicellular autotrophic organisms mostly composed of unicellular algae (microalgae) that freely float in the water and are transported by currents. There are several types of microalgae separated in five groups according to their shape and color : diatom, cyanobacteria, chlorophyceae, chrysophyceae, dinophyceae. Some algae can proliferate under favorable conditions (when temperature and /or nutrients such as phosphate and nitrate increase for example), causing blooms known as red-tides (Figure2). These phenomena are not always “red”, as the color of water is depending on the pigments of the microalga. Phytoplankton’s role as primary producer has an environmental impact on the food chain. As it uses solar energy directly as well as CO2 and mineral salts to produce organic matter and release oxygen, it is consumed by zooplankton. Zooplankton is mainly composed of tiny heterotrophic animals from marine surface. It is then consumed by fish or crustaceans (shellfish). 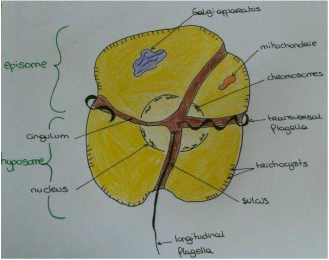 Figure3. Dinoflagellata morphology and main features Figure3. Dinoflagellata morphology and main features Dinoflagellate A dinoflagellate is a unicellular organism that can be autotroph thanks to endosymbiose with a microalgae. A dinoflagellate is characterized (Figure3) by the presence of two perpendicular flagella, the absence of histones in the nucleus, sub-cortical alveoli that most of the time contain cellulosic slabs regularly arranged. The host belongs to the class Dinophyceae (photosynthetic dinoflagellates). Dinophyceae have a dinokaryon : their chromosome remained condensed during interphase stage of mitosis (interval between two successive mitoses). The parasite belongs to class Syndiniophyceae (order Syndiniales, family Amoebophryidae). Syndiniales are, up to now, only composed of marine parasites. The taxonomy of this group is still uncertain, as most of Syndiniales are only known by their SSU (small sub-unit) rDNA gene sequences issued from marine environment. Amoebophrya is a dinoflagellate from MALV Group II (one of the V orders of Syndiniales[4], Marine Alveolates) which is only composed of parasitic dinoflagellates. How does the parasite Amoebophrya survive in the natural environment ? Why is the life cycle of dinoflagellates so important to understand host-parasite relationship and advantages ? All those questions remains and are waiting to be solved !
To be continued… [1] Ultrastructure Des Amoebophrydae. 1st ed. Jean et Monique Cachon, 1969. Print. [2] Images.google.fr. N.p., 2016. Web. 28 Aug. 2016. [3] Miller, John J., Charles F. Delwiche, and D. Wayne Coats. "Ultrastructure Of Amoebophrya Sp. And Its Changes During The Course Of Infection". Protist 163.5 (2012): 720-745. Web. [4] Guillou, L. et al. "Widespread Occurrence And Genetic Diversity Of Marine Parasitoids Belonging To Syndiniales ( Alveolata )". Environmental Microbiology 10.12 (2008): 3349-3365. Web. |