|
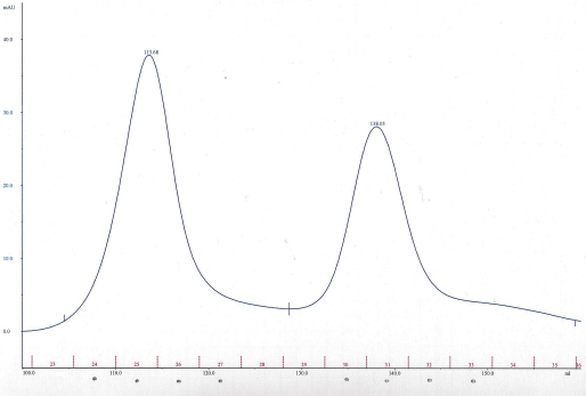
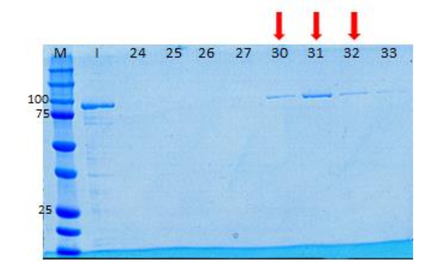
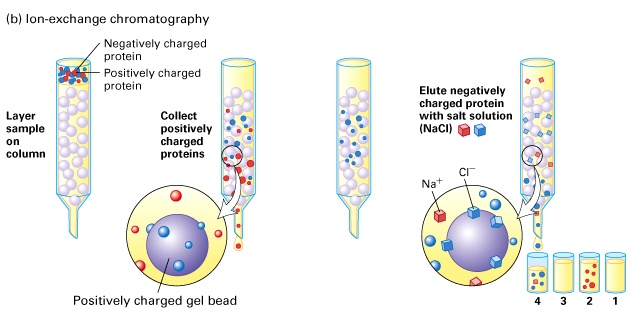
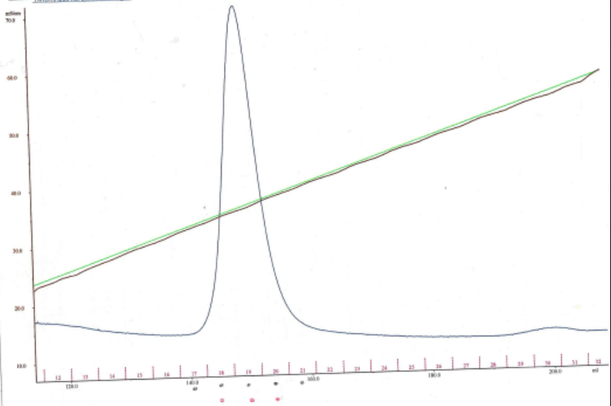
My aim then was to purify Hsp90 at a larger scale. To do so, I followed a six-step protocol. Here are the six steps : - Cell lysis - HisTrap affinity chromatography - Gel filtration chromatography - Anion exchange chromatography - ATP-agarose affinity chromatography - Storage of the purified protein It will be very long to discuss about all of these steps so I will simply talk about cell lysis, gel filtration chromatography, anion exchange chromatography and storage of the purified protein. Cell lysis This time, to lyse the cells I used sonication.  I pulled out my bacterial pellet from the freezer (-20°C) and I suspended it in a buffer solution. I placed my sample in an ice bath and I sonicated my bacterial suspension for 1 min then I paused for 1 min. I repeated this 4 times. Then, I centrifuged the obtained total lysate at 30000 rcf (relative centrifugal force) during 30 min. After that, I sampled the supernatant (soluble lysate) that I kept for the next steps. A sonicator. source : https://commons.wikimedia.org/wiki/File:Sonicator_togopic.png Gel filtration chromatography After HisTrap affinity chromatography, we mixed all the fractions that contained the protein and we got a final volume of 5mL. It's that sample of 5mL that I used for gel filtration chromatography. 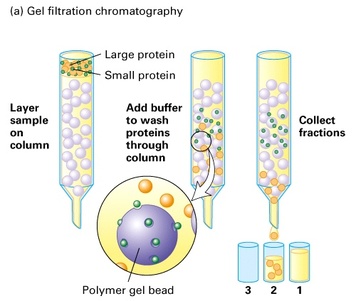 Gel filtration chromatography which is also called size exclusion chromatography separates molecules according to the size. Gel filtration resin can be thought as beads having little pores. Large proteins which cannot enter these pores will pass around the beads whereas small proteins will flow through the beads and thus take a longer path. The larger proteins will therefore elute first and the smallest ones will elute last. source : http://www.pha.jhu.edu/~ghzheng/old/webct/note1_1.htm Here is the graph that I obtained after gel filtration chromatography : The pics indicate the presence of proteins. To know if my protein of interest was collected in the fractions 24, 25, 26 and 27 (see black dots under the first pic) or in the fractions 30, 31, 32 and 33 (see black dots under the second pic), I did an SDS-PAGE gel. The presence of distinct bands around 90kDa (see red arrows) proved me that Hsp90 was in the fractions 30, 31 and 32. Anion exchange chromatography We mixed these three fractions and we got a final volume of 15mL. It's that sample of 15mL that I used for anion exchange chromatography. source : http://www.pha.jhu.edu/~ghzheng/old/webct/note1_1.htm In the anion exchange chromatography, negatively charged proteins will bound to the positively charged beads whereas positively charged proteins will pass through the column. Therefore, positively charged proteins will elute first. Negatively charged proteins are eluted with a salt solution. Here is the graph that I obtained after anion exchange chromatography : I collected the fractions 18, 19 and 20 (see red dots). After mixing these fractions, I got a final volume of 15mL. It's that sample of 15mL that I used for the next step. Storage of the purified protein
After ATP-agarose affinity chromatography, I concentrated the eluate and I obtained a volume around 180µL. After measuring the concentration of this aliquot of 180µL, we found that our protein was at a very low concentration. Indeed, we get six aliquots of 30µL concentrated at 1.23mg/mL. We kept these six aliquots at -80°C.
2 Comments
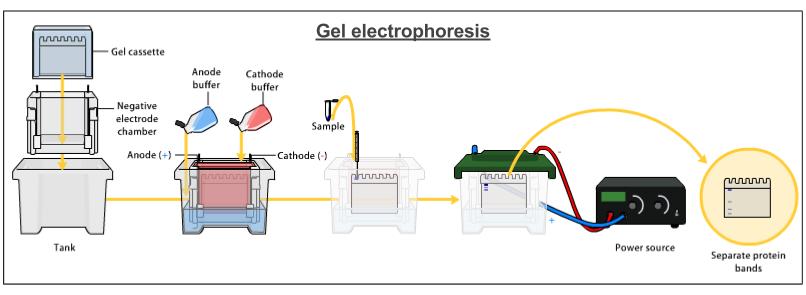
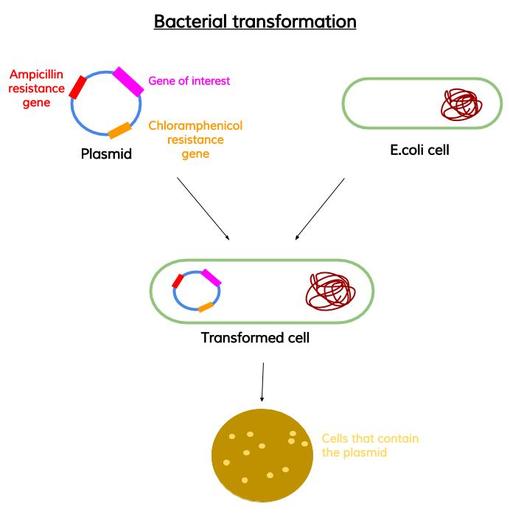
My aim then was to verify if the cells produced Hsp90 as intended. To do so, I made a recombinant protein expression test. There are three stages in this test : mini lysis of bacterial cells, mini purification by Ni-NTA affinity and SDS-PAGE analysis. 1° Mini lysis of bacterial cells The mini lysis of bacterial cells basically consists in five freezing-thawing cycles. I froze the cells in liquid nitrogen (around -210°C) then I thawed the cells at a room temperature water bath. The aim of this freeze-thaw method is to break cells so that they release cell components. By doing so, I obtained a total lysate which is composed of soluble and insoluble proteins, DNA, RNA, etc. I sampled 10µL of this total lysate for SDS-PAGE analysis. Remember, my goal is to keep only Hsp90 (which is a soluble protein) and get rid of the rest. Therefore, the next step was to separate soluble proteins from insoluble proteins and other insoluble components (like DNA). To do so, I centrifuged the total lysate and I sampled 10µL of the obtained soluble lysate (supernatant) for SDS-PAGE analysis. 2° Mini purification by Ni-NTA affinity To purify Hsp90 (which was previously tagged with histidine) I used Ni-NTA affinity chromatography. Indeed, this type of chromatography is used to purify histidine-tagged proteins. Next, I loaded the soluble lysate onto the matrix. His-tagged proteins will bound to nickel ions with high affinity. After washing, non His-tagged proteins will pass through the matrix. I then eluted the His-tagged proteins with a His buffer. I sampled 10µL of the eluate for SDS-PAGE analysis. 3° SDS-PAGE analysis I added loading buffer in each of my three samples (total lysate, soluble lysate and eluate) and I heated them at 95°C for 5 min. Then, I loaded the samples into the wells of an SDS-PAGE gel. I ran the electrophoresis at 180 V for 55 min. I stained my gel with Coomassie blue and here is what I got : 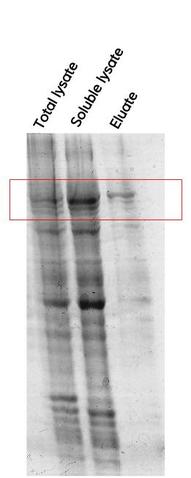 I could see, on the Eluate column, a distinct band around 90kDa which corresponds to Hsp90. However, on the Eluate column there were also other bands. It meant that my protein of interest wasn't completely pure. Therefore, the next step was to purify Hsp90 at a larger scale. As I told you in my previous post, Hsp90 is partly responsible for cancer but the best way to use Hsp90 inhibitors for anticancer therapy remains to be defined. Therefore, we need to keep studying Hsp90. It is obvious that to study Hsp90 we need to have Hsp90. But how can we obtain this protein ? The first thing to do is to transform bacterial cells so that they can produce Hsp90. After that, we can make a recombinant protein expression test that will allow us to determine if Hsp90 was produced by cells as intended. Then, we have to isolate this protein from other cell components, it's what we call protein purification. Today, I will simply talk about the first step which is bacterial transformation. My aim was to introduce into competent E.coli cells a pRSET plasmid which contained the gene coding for Hsp90 as well as an ampicillin resistance gene and a chloramphenicol resistance gene. To do so : - I mixed the pRSET plasmid with competent E.coli cells. - I let the mixture on ice for 45 min. - I heated the mixture at 42°C for 45 sec. - I placed back the heated mixture on ice for 5 min. - I added SOC media to the mixture. - I placed this bacterial suspension in a shaking incubator at 37°C for 45 min. - I plated some of the bacterial suspension onto an LB agar plate containing ampicillin and chloramphenicol. - I let the plate at 37°C overnight. source : me The presence of the antibiotic resistance genes on the pRSET plasmid allowed me to isolate bacteria which had the plasmid from bacteria which didn't have it.
- I inoculated few colonies from my LB agar plate. - I started a 20mL pre culture. - I then made a 100mL pre culture. - I then made two 1L cultures. - When OD = 0.8, I induced the expression of Hsp90 by adding IPTG* on both cultures. Indeed, at this OD, bacteria are in the exponential phase of growth, they are very active which is good to the protein production. - I centrifuged my cultures. - I discarded the supernatants. - I kept the pellets at -20°C. The next step will be to verify if the cells produced Hsp90 as intended. * Isopropyl β-D-1-thiogalactopyranoside These past two weeks, I tried to express a chaperone protein known as HSP90 (heat shock protein of 90 kDa). Production of proteins is a common laboratory technique because it gives a wide range of possibilities and especially in our case, its use to treat cancer. But what is the function of HSP90 and how is it related to cancer ?
Function of HSP90 What is a heat shock protein ? Heat shock proteins, as their name suggests, protect cells when they are stressed by elevated temperatures. Indeed, in response to stressful conditions (high temperatures) cells produce many heat shock proteins. Why do we say that HSP90 is a chaperone protein ? These heat shock proteins, once produced by cells, will play their role of chaperones : they will assist the folding of stress-denatured or newly synthesized proteins. Relation between HSP90 and cancer These proteins that depends on HSP90 for their maturation, are called HSP90 client proteins. Unfortunately, many of these client proteins control functions involved in malignant transformation. Thus, the guardian role of HSP90 is subverted into cancer. Therefore, finding HSP90 inhibitors seems to offer an anticancer strategy. However, the best way to use these inhibitors for anticancer therapy remains to be defined. |