|
I received an interesting question on my last blog post so I'd like to answer it.
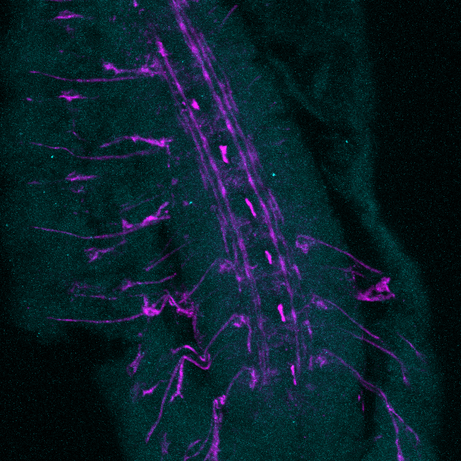
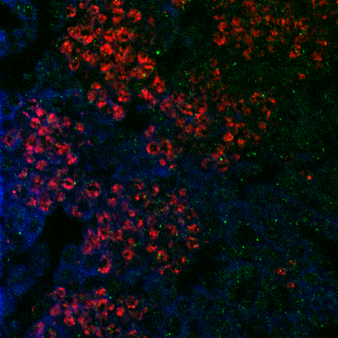
I was explaining that I had to increase the concentration of the primary antibody when doing my immunostaining to make sure that the protein I'm studying will be marked. I have been asked whether this could increase the background noise (which was already pretty high!). Well, I think this is a good point and usually the optimal dilution for the antibody is determined by running a serial dilution test. This should allow to find the best signal to noise ratio. If you look at the picture I got this morning, the background noise is effectively very high and there is no signal specific for the protein, which means that increasing the concentration of the primary antibody was not optimal.
5 Comments
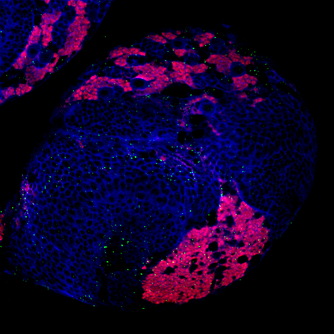
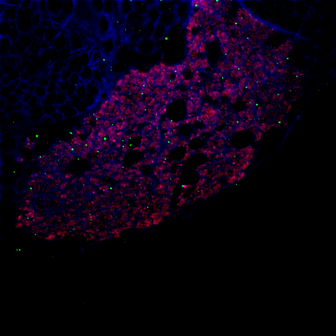
As you may know, I am trying to understand how Lola, a transcription factor expressed in early born neurons, can repress the expression of stem cell genes to prevent the neuron to neuroblast reversion. Lola interact with the Rcd5 protein which recruits the transcription machinery Previously, the lab I working on did a yeast two hybrid assay (you can read my blog post about it) and discovered that Lola might interact with Rcd5. Interestingly, Rcd5 has been shown to facilitate the recruitment of RNA Polymerase II preinitiation complexes (i.e. the transcription machinery) to the promoter regions of target genes (Anderson et al, 2010). The protein promotes genes transcription required for normal proliferation. Localizing the Rcd5 protein using immunostaining and confocal microscopy In order to localize the Rcd5 protein in the Optic Lobe, where Lola is required to maintain immature neurons differentiated, I stained wild type Drosophila larve brains. I used mouse the anti-Dlg antibody because Dlg is a marker of membranes and should indicate the shape of the brain. It is stained in blue in the figure. Rat anti-Elav is stained in red. Elav is a transcription factor specifically expressed in early-born neurons. Finally, guinea pig anti-Rcd5 (obtained from Anderson’s lab) stained in green shows the protein I am interested in localizing. I used a confocal microscope to obtain images, with a 40x and 63x magnification. As you can see, it is not clear that Rcd5 (in green) is preferentially expressed in the neuroblasts (big dark dots). What’s the best between staining larve brain and embryo? So I repeated the staining in embryo of wild type Drosophila, because immunostaining are usually more efficient than in larve brains. I also tried different concentration for the anti-Rcd5 antibody: 1:500 because Anderson et al used this concentration and a higher concentration of 1:100. However, this did not give more conclusive results and it is not clear where the protein is localized. Reliable staining or Residual fluorescence?
Because the staining of the protein (brightness of the color) is so weak, I was wondering whether the green dots that I was observing on the picture could only be some background noise from the fluorophore. Indeed, to perform an immunostaining, a primary antibodies is used to target the protein of interest, in my case Rcd5, and a secondary antibody is used later in the process to add fluorescence. So, the green dots on the figures might only be some residual staining from the secondary antibody instead of the actual protein. This would be possible because the primary antibody, anti-Rcd5, is quite ancient now (from 2010). To test whether the anti-Rcd5 antibody is efficiently marking the Rcd5 protein, I decided to:
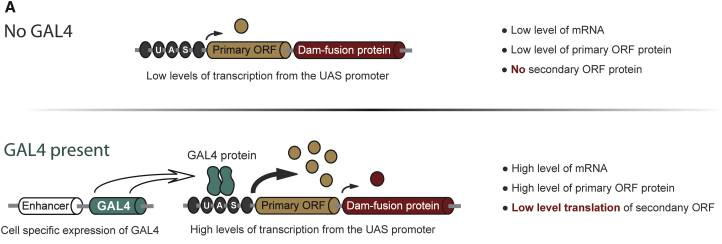
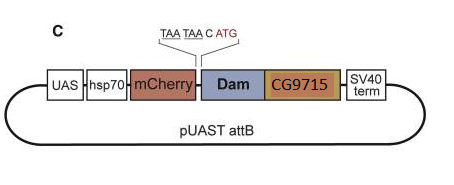
Although my slides are ready, I’ll have to wait until tomorrow to use the confocal microscope and know the answer… Reference: Andersen, D.S., Raja, S.J., Colombani, J., Shaw, R.L., et al. (2010) Drosophila MCRS2 associates with RNA polymerase II complexes to regulate transcription. Molecular and cellular biology. [Online] 30 (19), 4744–4755. Available from: doi:10.1128/MCB.01586-09. I am doing my internship in a lab which has developed a technique for identifying the transcription factors expressed in a cell-specific manner. In other words, it gives information about the genes being transcribed in the cell-type you are studying, such as neurons or neuroblasts, at different stage of development, such as embryonic or adult stages. This is very interesting because as I explained in my first blog post, stem cells differentiate to specific fate following a transcriptional program of the genome. Unlike other existing methods, this technique called TaDa can be used in vivo and does not require cell purification. Methylation as a tag to identify the genetic targets of a protein More generally speaking, the method allows us to identify the genes which are bound by a protein of interest for any model organism used (and the while genome is screened). This is achieved through the fusion of an enzyme, DNA adenine-methyltransferase from Escherichia coli, to the protein of interest. The genes with which the protein of interest interacts will be identified and tagged by a methyl group, attached by the enzyme, to the adenine base of the DNA. The fusion protein is placed under the control of UAS activation system so that tissue-specific expression can be achieved using the GAL4 promoter (GAL4 is placed under the control of promoter naturally expressed in neurons, neuroblasts or other specific cells). You can find more information about the GAL4/UAS system in my previous blog post. Methylation: a toxic tag? However, Dam methylase must be expressed at very low level because high methylation of DNA can be toxic for the organism or nonspecific. The issue at this point was thus to reduce levels of Dam methylase translation and avoid toxicity. Because ribosomes can reinitiate transcription, the lab split the construct with a codon stop. This codon is located after the UAS and the primary Opening Reading Frames to reinitiate and reduce levels of translation of the Dam-fusion protein (see below on fig.2). Hence, on the one hand, the primary ORF (ORF1 in the drawing) is translated at high levels. This is interesting as it encodes a fluorophore, mCherry, allowing us to locate the construct within cells. On the other hand, the secondary ORF encoding Dam methylase is supposed to be translated at low levels after reinitialization of translation (Fig.1). Without GAL4 being expressed, the primary ORF only would be translated at low level. So, the genetic targets of the protein of interest will be tagged in a cell-type specific manner. The lab demonstrated that this construct was not toxic at any stage of development. Is the technique accurate? They have assessed the accuracy of their new technique using RNA Polymerase II as protein of interest as it has been studied many time in previous studies. TaDa technique accurately identified RNA Pol II occupancy in the genome when compared with other methods! I will use this technique! One of the goal of my project is to use this technique to determine the genetic targets of the Zcchc7 protein, involved in the nuclear exosome and interacting with Lola. I am thus constructing a plasmid where Dam is fused to Zcchc7 so that the DNA bound by Zcchc7 will be tagged by methylation (Fig.2)! Reference
Southall, T.D., Gold, K.S., Egger, B., Davidson, C.M., et al. (2013) Cell-type-specific profiling of gene expression and chromatin binding without cell isolation: Assaying RNA pol II occupancy in neural stem cells. Developmental Cell. [Online] 26 (1), 101–112. Available from: doi:10.1016/j.devcel.2013.05.020. I am sure you can all remember this biology course in high school when you learn how to cross the flies Drosophila to get curly wings, red eyes… In this blog post you will discover how these considerations can be used for research. GAL4/UAS system The GAL4/UAS system is widely use to direct the expression of gene sequence of interest in specific tissues (1). GAL4 is a yeast transcription factor yeast which can activate gene transcription by binding to an Upstream Activation System (UAS). In labs working with flies, the idea is to use two lines: - A driver lines expressing GAL4 in a tissue specific manner - A responder line expressing the gene of interest in every tissue but this coding sequence is under the control of UAS site. After crossing, the transcription of the gene of interest only occurs in the cells expressing both UAS and GAL4. GAL4: tissue-specific expression You may wonder how GAL4 can be express specifically in tissue. The identification of transcription factors expressed in specific cell population are used as cell-type “marker”. For example, Elav is a transcription factor specifically expressed in neurons or Insc in neuronblasts. Thus, injecting Elav-GAL4 or Insc-GAL4 in a fly lead to the expression of GAL4 in neurons, or neuroblast respectively. Crossing flies: expression of GAL4 and UAS simultaneously My goal is to downregulate the encoding sequence of Zcchc7 (CG9715) using short hairpin RNA (shmiR) to assess the function of the protein in the Central Nervous System. (figure 1)
Balancing chromosome to keep track of the UAS construct in flies’ genome As Elav and Insc are natural transcription factor, GAL4 will naturally be expressed. However, it is not that simple for UAS-CG9715-shmiR sequence. Indeed, this has been injected into flies and we need to keep track of the sequence as it could undergo chromosomal recombination. To identify the flies that integrated the construct in their genome, one chromosome over the two of the pair is used as a balancing chromosome. And here you’ll see the power of curly wings. The balancing chromosome does not express any functional genes except Cy0 encoding curly wings. Therefore, flies with a pair of balancing chromosome will express any functional genes from except curly wings, and will die. However, those expressing one balancing chromosome and on the other chromosome of the pair will express be healthy (functional genes on one chromosomes) and have curly wings! Flies with straight wings have a pair of chromosome with UAS-CG9715-shmiR and any balancing chromosome. On the figure you can see the possible outcome of crossing involving each of these phenotype. Reference 1. Duffy, Joseph B. "GAL4 System Indrosophila: A Fly Geneticist's Swiss Army Knife". genesis 34.1-2 (2002): 1-15. Web. For those who prefer watching video than reading, the beginning of the one below is for you! In 2014, my supervisor’s lab identified a transcription factor which is required to prevent dedifferentiation of newly formed neurons (1). These immature neurons revert to neural stem cell characteristics if they don’t express Lola. How can Lola fulfil this unknown biological function?
I will focus my investigations toward the study of two molecular interactions using Drosophila as a model organism
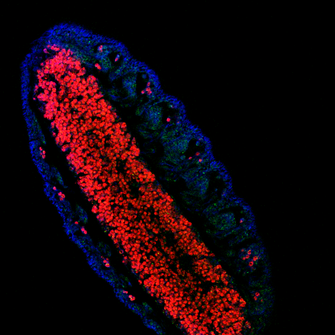
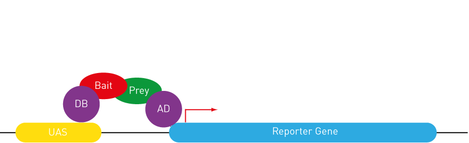
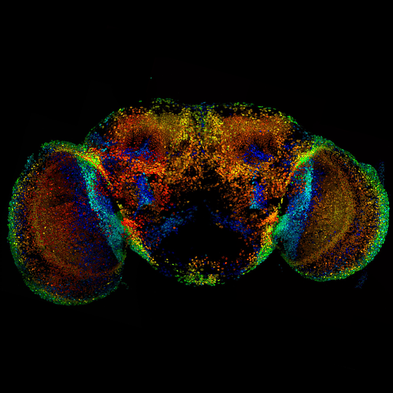
understanding the function of lola for newly formed neurons maintenance: how flies can help (Cécile)18/3/2017 Lola is a transcription factor required to maintain newly formed neurons in a differentiated state and prevent them from reverting to the neural stem cell type initiating tumors. This function implies the repression of neural stem cell genes and the activation of other genes’ transcription. So, Lola must interact with other factors to ensure opposite functions. Identification of potential co-factor for Lola The yeast two hybrid assay is very useful to determine protein-protein or protein-DNA interactions, by tracking the resulting downstream effects. The reporter gene chosen must encode a measurable transcription product, which is produced if a functional GAL4 binds to the Upstream Activation System (UAS). The principle of the assay is to split GAL4 in two fragments and attach a protein of interest to each. The first one contains the binding domain (BD) recognizing the UAS sequence. The second one contains the activating domain (AD) initiating transcription. Transcription can only be induced if the both fragments are recruited. In Lola’s case, Lola was used as a Bait, attached to GAL4 BD. A large number proteins from a drosophila DNA library was tested as Prey, attached to GAL4 AD. When the transcription product was observed, the protein from the library was considered as a candidate co-factor for Lola. Lola can interact with a protein recruiting the transcription machinery Rcd5 was detected through the yeast two hybrid assay. This protein is involved in the regulation of gene transcription by facilitating the recruitment of RNA Polymerase II Complex to the promoter region of target genes (1). Understanding how Lola fulfill its function could therefore benefit from investigating its interaction with Rcd5. As any previous research has been done, I will first practice a staining of Rcd5 using fluorescent antibodies within adult Wild Type (1118) Drosophila brains. Although the distribution of this essential protein is expected to be ubiquitous over the brain, I will maybe observe up-regulation in specific cell type… Following this general overview, I will use a Drosophila line over-expressing Rcd5 to get more information about protein function, although the precise experiments has not been designed yet. A protein involved in degradation of RNA can interact with Lola, to modulate its function? The yeast two hybrid assay has revealed many other proteins likely to interact with Lola. As I explained it to you on my previous blog post, removal of Lola only causes dedifferentiation of immature neurons. It means that more permanent mechanisms are recruited to repress neural stem cells genes throughout development. So, some factor might bind to Lola to modulate its function. I will be investigating one of them, the Zcchc7 protein, which appeared to be involved in the nuclear exosome. This complex acts as a quality-control system that degrades non-functional RNA but also regulate normal mRNAs and repress viral RNA in the nucleus (2). Nothing is known about its interaction with Lola and Zcchc7 could both enhance and repress expression. I began to investigate the role of this interaction using a drosophila line in which the sequence encoding this protein were down-regulated by RNAi in the Nervous System only. However, staining adult brains with antibodies did not reveal any tumours or abnormal brain development. This might be due to the low efficiency and lack of specificity of RNAi. In any case, antibodies staining gives amazing pictures, and I wanted to show you one from my lab, there are few other on the website… Crossing flies to direct the expression of target genes
I am also setting up a crossing. In Females genotype, GAL4 protein encoding sequence is placed under the control of promoters naturally expressed in neurons and neuroblast only. In Male genotype, a small hairpin RNA targeting Zcchc7 encoding sequence has been inserted. As expression of this interfering RNA requires GAL4 to bind the UAS, Zcchc7 downregulation will only occur in neurons and neuroblast in the progeny for which I will conduct a survival assay throughout embryos development. I will maybe try to stain some adult brains to compare this way of silencing gene expression with my previous results. Insert genes of interest into flies: construct the plasmids first Finally, I am constructing 3 plasmids that will be infiltrated into flies. The first contains a GFP tag to localize the Zcchc7 encoding sequence within the tissue. The second one is a control without GFP to ensure that the tag does not affect drosophila physiology. The last one contains a fusion protein: once flies are transfected, the sequence encoding Zcchc7 protein also encodes Dam methylase. This enzyme methylate at low level the molecular targets of the protein! The technique is called TaDa and has been set up by the lab I am working on itself (3). However, I have to deal with several issues while trying to make the plasmids, mainly due to the large size of the insert (5 kB). References
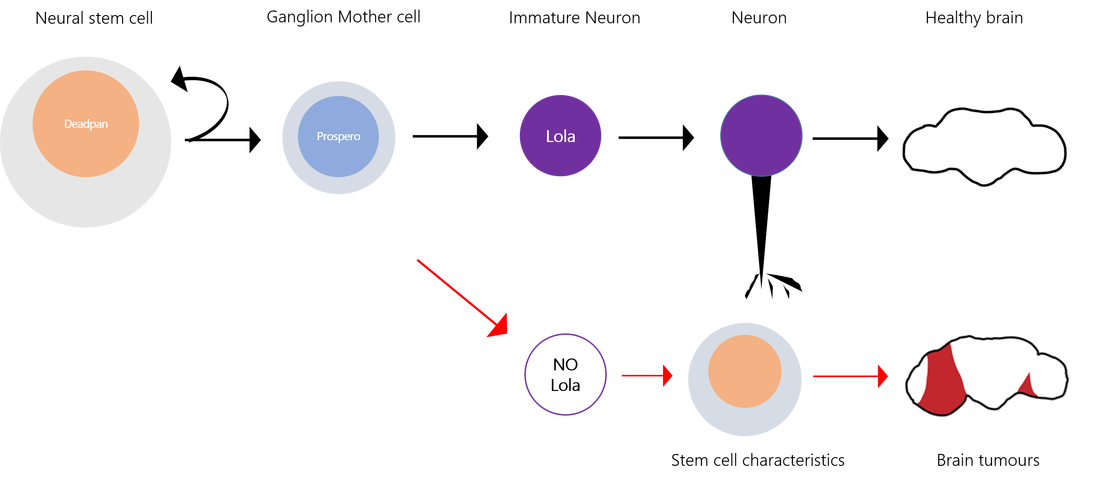
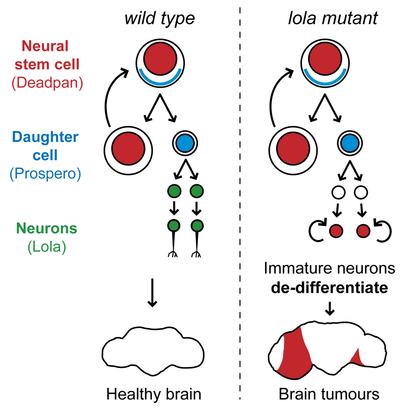
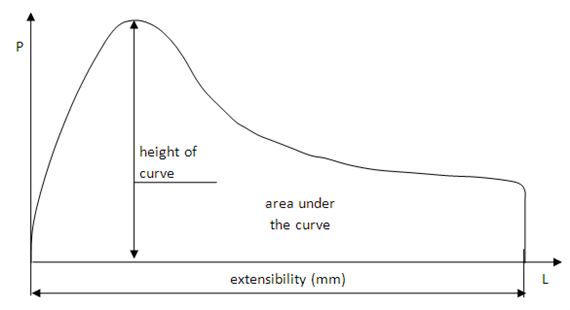
Complex organisms as we are rely on the differentiation of stem cell to form muscle, blood, immune cells and... neurons. During nervous system development, neural stem cells asymmetrically divide and give rise to two daughter cells. A new neural stem cells ensures self-renewal. Ganglion Mother cells (GCM) then further differentiate into neurons and glial cells to fulfill specialized functions. Which factor control the choice between neural stem cell renewal and differentiation? Stem cells are carrying the same genetic information but the regulation of gene expression determines the outcome of a division. This highly ordered activation of repression of genes results in modifications of cell’s shape, size and metabolic activity. The Southall lab , where I am currently doing my internship, has investigated the factors determining the fate of neural stem cells, using Drosophila melanogaster as a model organism (reference for the original research paper below). The transcription factor Deadpan is expressed in the nucleus of initial neural stem cell to drive cell division. At this stage, another TF, Prospero, is expressed in the cytosol of one side of the cell. If it enters the nucleus, it represses genes required for self-renewal and activates genes for neuronal and glial cell differentiation. So, Prospero determines the choice between self-renewal and differentiation. In prospero mutant, GMC effectively failed to differentiate and they reverted to stem cells characteristics. Deadpan is not repressed anymore, cells start dividing, and eventually tumors can be initiated. Because Prospero appeared to both activate and repress gene expression, it was likely that some co-factor was collaborating to modulate its activity. The sequence which is bound by Prospero binding has been extracted and many other proteins which recognized this motif were identified. However, a yeast one hybrid assay, by which DNA-protein interaction can be assessed, has revealed Lola-N, a specific isoform from the 25 splice variants of the BTB-zinc finger transcription factor Lola. So, Lola, a transcription factor, seems to bind the same molecular targets as Prospero. Are they acting together? When and where is Lola expressed? A widespread method used to answer this question is the staining with antibodies, which label proteins with fluorescent tags. Lola appeared to be expressed in post-mitotic neuron, after GMC division and Prospero expression. So, the brief expression overlap between Lola and Prospero does not ague for their collaboration. How does Lola act in postmitotic neuron? Lola's binding site and interactions with its targets were identified. Lola represses several neural stem cells genes such as deadpan and other genes initiating cell cycle. That is why it is called a tumor suppressor. However, Prospero can repress neural stem cells genes as well! Because Lola expression level only increases at the final division of GMC and then in post mitotic neurons, it is likely that Prospero initiates cell differentiation while Lola is required to maintain newly formed neuron in a differentiated state. As expected, in lola mutant, removing Lola resulted in the dedifferentiation of immature neurons, which began to divide and form tumors. Therefore, although it was well-known that cells could be reprogrammed and re-differentiated into the desired lineage, these results provided new information on factors required to prevent dedifferentiation. On the other hand, Lola removal does not cause dedifferentiation of mature neurons… More permanent mechanisms might be recruited later in the development. So, something might bind Lola to regulate its function… My aim for this project is to investigate one of the factor likely regulating Lola’s function and I’ll explain how in my next blog post... A perspective for regenerative medicine? Lola prevents immature neurons from dedifferentiating … imagine if we could induce neurons to dedifferentiate, and then direct their differentiation to neurons of our choice, it would be an amazing method of repair of diseased tissues in vivo! Even more since the same factor can maintain repression of neural stem cells genes, avoiding cell division and gliomas… Useful links: Southall, T., Davidson, C., Miller, C. & Carr, A. et al. (2014) Dedifferentiation of Neurons Precedes Tumor Formation in lola Mutants. Developmental Cell. [Online] 28 (6), 685-696. Available from: doi:10.1016/j.devcel.2014.01.030. Only 1 minute to understand neural stem cells? Watch this video! If wheat quality vary each year because of weather, industry and bakeries must be able to sell product of a constant quality over years. It is thus a challenge for mill to get informations on flour quality and determine the best usage for them. A widely used technique for this is alveograph. The principle is to inflate a sheet of dough and measure how much pressure and how much time is necessary to create and burst an air bubble in the dough. Look at the principle on this great video created by ChopinTechnologies Step 1: make a dough Make kind of a bread dough by hydrating flour at 50% with salted water (25 g per liter) and knead for 8 minutes at 24°C. After a 15 minutes break, the circular dough is deposited on the alveograph and homogeneously spread into a thin sheet. Step 2: Forming bubble Then, the air blast starts inflating dough and forming the bubble. Simultaneously, a computer is collecting and drawing a curve (fig 1). Fig 1: Curve from Alveograph [2] Step 3: Analyse the curve
Thus, the bubble gets bigger with less pressure necessary, until burst. The height of the bubble (in mm) that was achieved is called L (extensibility on fig 1). The ratio P/L is an interesting information. P/L > 1: a high pressure is necessary so we have a strong dough. However, L is small so the bubble burst when still small. That kind of flour can be used to reinforce other, and we call it an improving wheat. P/L < 1: a small pressure is necessary so dough is weak but as L is high you can make big bubble. Concretely it correspond to biscuit flour, around 0,6 or 0,7, which do not need to inflate as for bread. Thus P/L ratio give a good idea of flour best usage.
These two information complement each other to determine flour quality. Let’s take the exemple of the P/L ratio of 0,7 again: although it correspond to biscuit flour, if you obtain a high W then flour is still adapted to bread making. Usually, W > 300 mean a strong flour, ideal for bread.
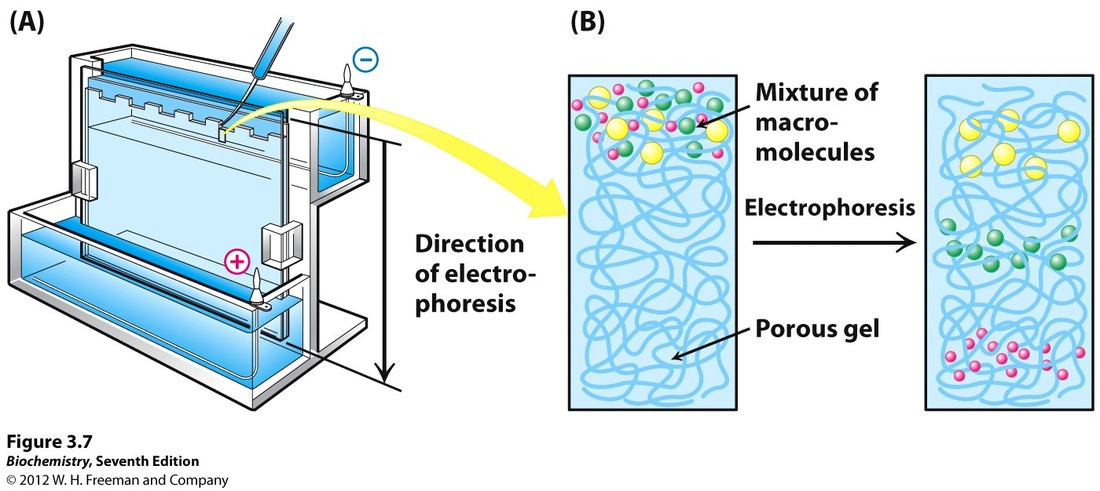
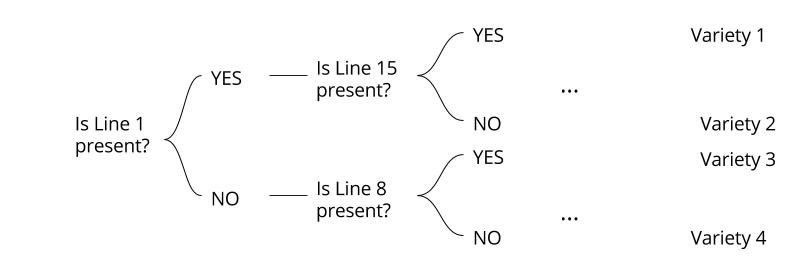
For example to make croissant and batter dough, you need a flour with a high P/L ratio strong dough but still elastic (high Ie). References [1] Oulton, Randal. "Alveograph". CooksInfo.com (2005): n. pag. Web. 25 Sept. 2016. [2] "Quality Analysis Of Agricultural Products|Digital Textbook Library". Tankonyvtar.hu. N.p., 2016. Web. 26 Sept. 2016. electrophoresis, a way to identify A wheat grain variety by the migration of its protein (Cécile)26/9/2016 Poly-acrylamide gel electrophoresis (PAGE) is a technique well known in laboratory of biotechnology or biochemistry. I don’t know this technique… reminder: A mix of macromolecules, protein or nucleic acids from DNA molecules, is submitted to an electric field (fig 1). Thus, charged molecules migrate on gel. This migration occurs more or less quickly depending on several parameters such as length, conformation and charge of molecules. It allows to separate (fig 1 (B)) the different molecules in the initial mixture and to deduce information like molecular mass… Fig 1: Principle of electrophoresis  A concrete application of this scientific method is the identification of wheat varieties, and you’ll discover in this article how. Proteins of wheat are spread into 4 classes: albumin, globulin but also gliadin and glutenin, wheat storage proteins. Gliadin can be separated by electrophoresis. [1] The great diversity of gliadin (around 120 alleles) [2] generate specific profiles for each seed variety. Selection 40 representative grains from billions of seeds Some wheat contracts are appropriated to bread making industry but some other are only suitable for animal feed. So let’s imagine that you are working in a company that sells flour for milling activities and you receive something like 30 trucks a day full of wheat. How to be sure that your supplier is really delivering milling wheat ? You can determine one seed variety using PAGE. But you should wonder how to do when you receive several tonnes, corresponding to several billions of seeds of wheat a day? The idea is make the analysis on a representative sample taken every 300 to 500 tonnes delivered. A representative sample initially weighs 1 kg and then an apparatus is used to remove 1 seed every 10 seed. You obtain 40 representative grains from billions of delivered grain. Prepare a mixture of macromolecule for each seed First of all you need to prepare the mixture that will be used for electrophoresis. Each seed has to be crushed and put in a solution of ethanol chlorure, saccharose and crystallised purple dye for 24 hours. The day after, the mixture is stirred for 15 minutes and centrifuged for 5 minutes. It is ready to be deposited on gel and start electrophoretic migration. However, at this end of this step, it is impossible to see the seed profile, and just like pictures needed to be developed, we use a solution of electrophoretic profile is soaked on trichloroacetic acid and coomassie blue dye to bring out the lines. Indeed, coomassie blue binds to amino acids (lysine and arginine mostly) of proteins The longer you let it soak, the strongest the colour will be. Usually, it is let 24h. From a column of blue line to the name of wheat variety The 40 profiles appear on 40 columns on the analytic paper (fig 2) from electrophoresis, each one corresponding to one seed. A profile is constituted by several blue lines numbered, corresponding to gliadin (or glutenin) fractions. Several years of observation and study of profiles from wheat of a known variety allowed to create trees or key of identification. Fig 2: analytic paper : electrophoretic profiles Thus, line by line, you follow a branch leading to the name of the seed variety (fig 3). Fig 3: Tree of identification What is the electrophoretic profile of milling wheat?
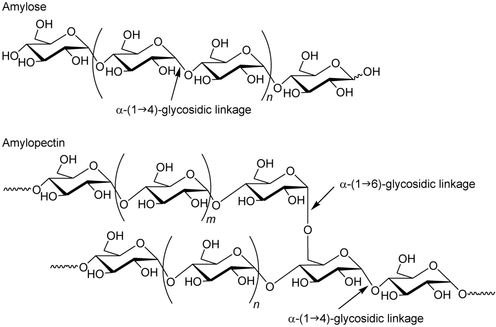
The difficulty is that a milling wheat contract can be a mix of several varieties. So you’ll not obtain 40 times the same profile, but something like 10 to 15 different profiles. Results are expressed in % of a variety. If 7 seeds or more among the 40 (15% of the contract) are of a non milling variety, the whole contract is considered as non compliant. Determining Wheat geographical provenance As all French variety has a known key of identification, this technique can also be used to determine wheat geographical provenance, an important information for company promoting “French product” for example. PAGE, a perfect technique? However, this technique pose certain problems. Proteins from damaged seeds can be not visible because they don’t bind to dye or need too much time to take a sufficient coloration. This technique is long: if you counted well, 3 days are necessary between sampling and variety identification. Any solution? A faster method, PCR (polymerase chain reaction), can also be used to amplify and study DNA from a single seed but it can be expansive and more complicated to practice and analyse. References [1] "Différenciation Des Variétés De Céréales Par Électrophorèse". Agrarforschungschweiz.ch. N.p., 2016. Web. 25 Sept. 2016. [2] KLEIJER, G. Sélection Des Variétés De Blé Pour La Qualité Boulangère. 1st ed. Station fédérale de recherches en production végétale de Changins, 2016. Web. 25 Sept. 2016. Making good bread require, of course, good bakers but also… good wheat! Indeed, you cannot use the same wheat to make flour for toast, sandwich bread, brioche, baguette… In order to decide of a suitable use for a wheat contracts, some tests are practised in agriculture field laboratories. Starch is the major component of flour as it represents at least two thirds of the weight of grain [1]. As nature does things well, a enzyme necessary for bread making industry is naturally present in wheat seeds: it is alpha-amylase. This enzyme is able to transform starch into simple sugar molecules, glucose and maltose. Focus on Alpha-amylase: from starch to maltose and glucose [2] Starch is a mix of two polysaccharides, amylose and amylopectin, at different rate depending on the plants. This molecules are special because there are composed by a same building-block: glucose. That is why there are called homopolymers.
Figure 1: Amylose and Amylopectin In both case, these oxygen bond can be broken by adding water molecules. One units gets an OH- from the water molecules and the other part gets an H+ to make the OH at carbon 1 and carbon 4. Alpha-amylase hydrolyses polysaccharides of starch using this process. The bond breaks can generate free glucose or pairs of glucose, that is to say maltose. A good rate of maltose for a good dough fermentation Maltose formation rate from starch determine dough fermentation depending on alpha-amylase activity. . Without enough maltose, fermentation is limited so breads are flat and moist test is not well developed. However, too much maltose leads to sticky and dough weak in consistency. [3] When does it happen? A persistent rainy whether before or during harvest can lead to the undesirable germination of mature wheat seeds. Indeed, alpha-amylase rapidly attack wet starch. [4] We call this the sprout damage, which would be disastrous for bread-making quality! Why? Because damaged starch is able to absorb more water and consequently it gives sticky dough. [5] So alpha-amylase levels mustn't be too high in order to limit starch degradation. Hagberg and the falling number method or how to determine alpha-amylase activity At the end of the 1950s, Sven Hagberg and his co-worker Harald Perten precisely developed a method, called the Falling Number method, to determine the rate and activity of amylase in wheat flour. Thus, it gives informations about a possible sprout damage. [4] As this test can be practised on grain silo intake, it allows to choose the best wheat contracts to sell for bread making. A specific apparatus is needed to practice the falling number test. Figure 2: Hagberg apparatus Distilled water is added to flour at ambient temperature. However, starch is not soluble to cold water. So after a vigorous shake insuring a homogeneous mix, this test tube is placed in a boiling water bath. The operator stir the sample during 60 seconds. Starch begins to gelatinize under the increase of temperature so the slurry becomes more viscous. But the high temperature also promotes starch breakdown into glucose and maltose by alpha-amylase contained in the flour. As a consequence, we observe a decrease of the slurry viscosity. Recording slurry consistency levels allows to know alpha amylase activity. The viscosity is measured as the time (in seconds) necessary for the stem to fall into the test tube. Thus, the higher the activity of the alpha-amylase, the lower the viscosity will be and the lower the falling number will be. So how to know if a wheat contract is a good one for bread making? [5]
Any solutions?
It is easy to improve high falling number wheat contract by adding malt (germinated barley seeds) but for small falling number, the only solution is to mix wheat with a wheat of better quality. [6] References [1] Analyse Des Farines. 1st ed. La m eunerie Milanaise. Web. 15 Sept. 2016. [2] "Alpha Amylase". Science.marshall.edu. N.p., 2016. Web. 15 Sept. 2016. [3] "Hydrolases #Boulangerie.Net". Boulangerie.Net. N.p., 2016. Web. 15 Sept. 2016. [4] "Falling Number". Wikipedia. N.p., 2016. Web. 15 Sept. 2016. [5] [email protected], LEMON. "Analyse Du Temps De Chute Hagberg". Ssl10.ovh.net. N.p., 2016. Web. 15 Sept. 2016. [6] Lachance, Pierre. L’Indice De Chute (Hagberg). 1st ed. 2004. Print. |