|
Some details of the process by which a hematopoietic stem cell becomes a mature immune cell remain unclear to this day. To tackle this issue, a new approach that already brought lots of new information is used: studying hematopoiesis at a single-cell level. It is to that extent that the project of my internship aims at developing a new tool to study single-cell dynamics in hematopoiesis. To do this, we design a new cell staining method using multiple combinations of proliferation dyes. It allows to follow the differentiation of 44 single hematopoietic stem cells as well in vitro as in vivo.
0 Comments
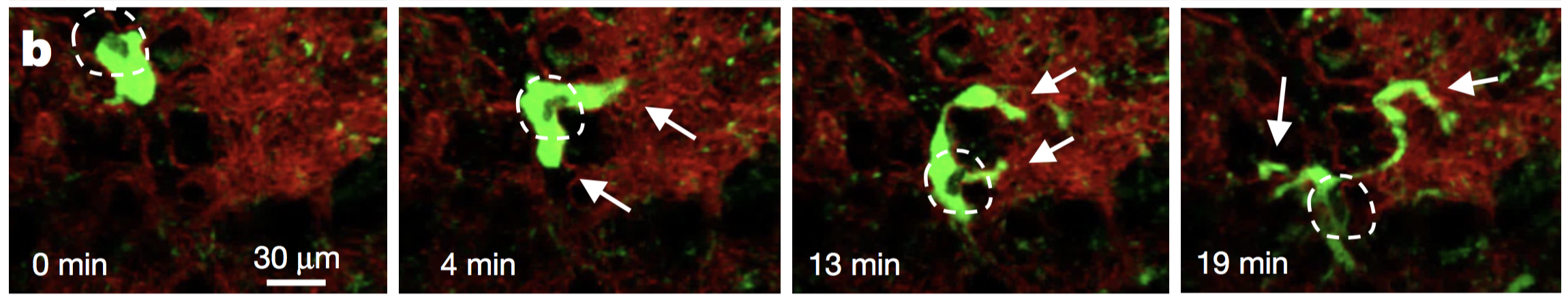
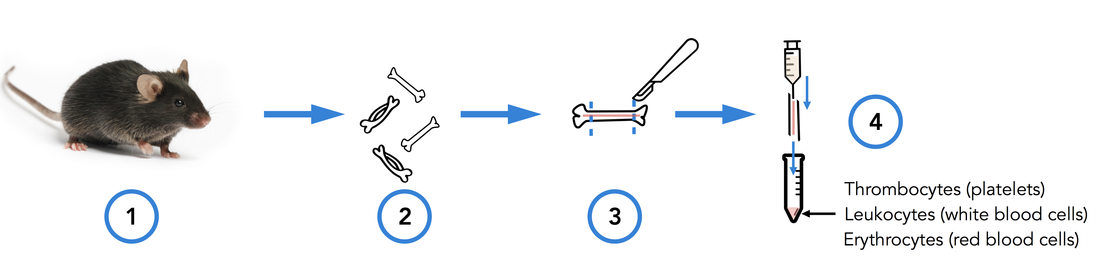
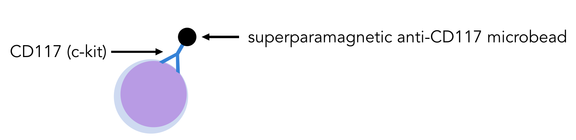
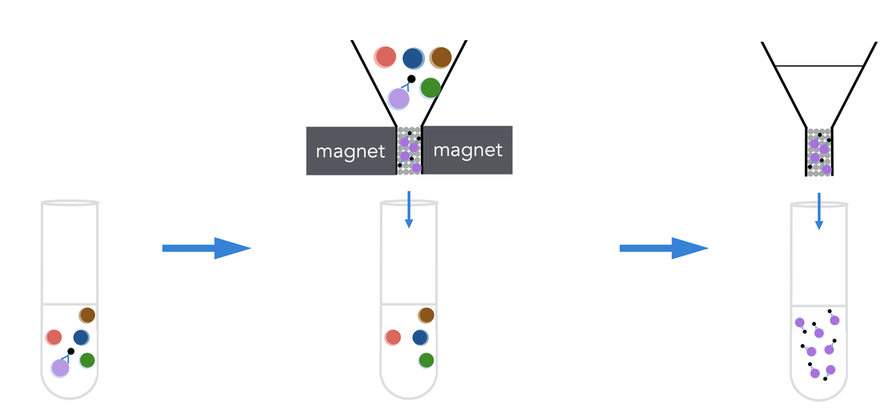
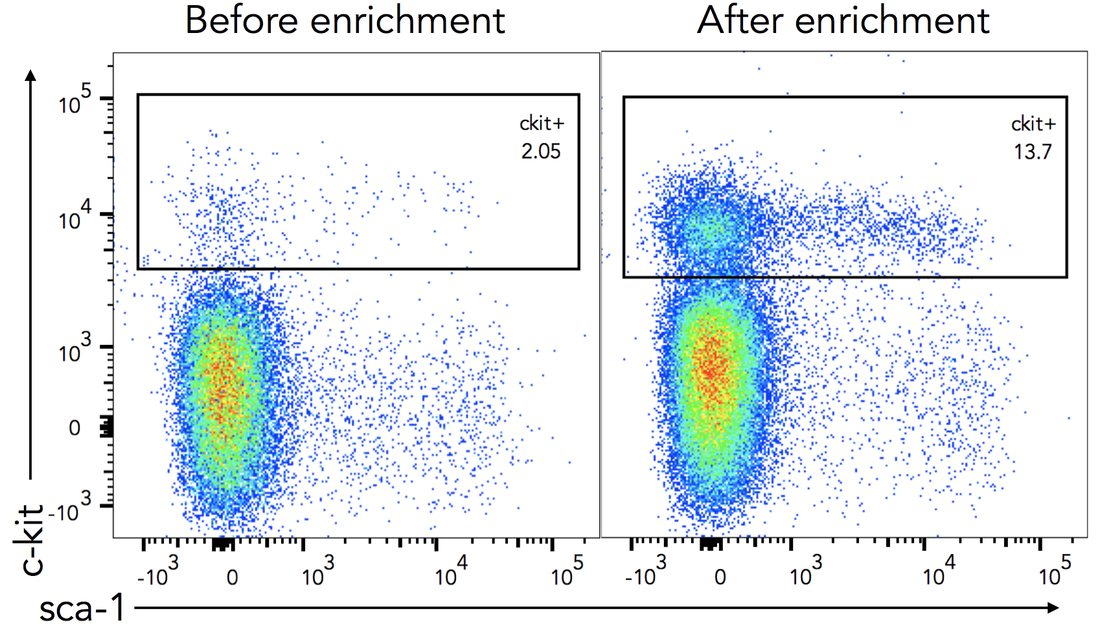
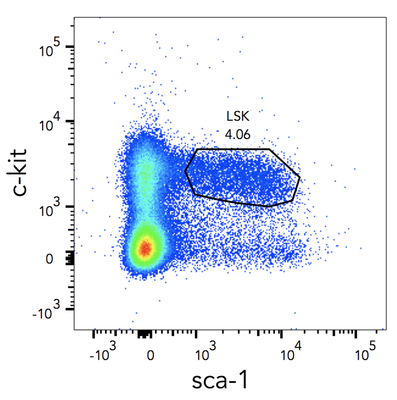
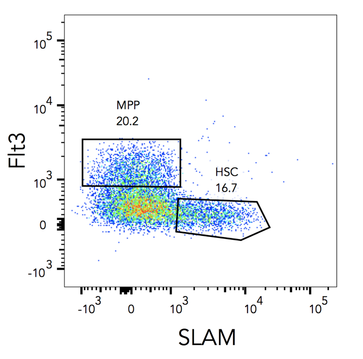
I wanted to share my enthusiasm with you in this blogpost, by telling you about an article that was published online in Nature two weeks ago. Researchers from Universities of California San Francisco (UCSF) and Los Angeles (UCLA) just discovered the lungs produce more than 50% of mouse platelets. What’s more, this organ has been revealed to host a reservoir for hematopoietic progenitors, capable to restore blood production in case of hematopoietic stem cell deficiency in the bone marrow. This finding was made possible by an imaging technique called two-photon intravital microscopy (2PIVM) allowing to visualize individual cells’ behavior in a mouse lung. They used a genetically engineered mouse strain that expresses GFP in megakaryocytes (platelet-producing cells) and platelets and tomato mCherry in all other types of cells. Here you can see a megakaryocyte (green) in the lung capillaries (red) of the mouse, undergoing platelet formation (arrows). Megakaryocytes had already been observed in the lungs but they were thought to produce platelets in the bone marrow, where hematopoiesis occurs. Video microscopy also revealed the presence of some megakaryocyte progenitor cells and HSCs in the lungs. To understand how they move between bone marrow and lungs, they did a set of different transplants. For instance, the researchers transplanted healthy donor lungs with fluorescent cells into a mutant mouse having an HSC deficiency. They soon detected fluorescent cells stemming from the donor lung into the recipient bone marrow. This shows that the reservoir of progenitors cells of the lungs can migrate to the bone marrow to repair bone marrow defects ! This can have a huge impact to treat blood diseases such as thrombocytopenia (a very low count of platelets), but gives also new perspectives to study stem cells. It seems essential to understand how the lungs and bone marrow collaborate to produce healthy blood cells. The article: E. Lefrançais, G. Ortiz-Muñoz, M. Looney et al. “The lung is a site of platelet biogenesis and a reservoir for haematopoietic progenitors”. Nature, 2017; DOI: 10.1038/nature21706 As I told you in my last post, I abandoned my dear Jurkat cells and started to work with hematopoietic stem cells (HSCs). Here I will show you how to isolate them from a mouse. 1. Get the cells out of the bones Bone marrow is the site of HSC’s biogenesis and thus contains most of them. It's also hosting a lot of other cell types, such as progenitors and mature immune cells. That’s why we have to get bones in order to find some hematopoietic stem cells. So first, we sacrifice a mouse (1). Then we take out its femurs, tibias and iliac crests (2). We cut both sides of the bones (3) so that a needle can pass inside. Then we flush the bones (4). This gives us all the types of cells the bone marrow contains: HSCs and progenitors, but also red blood cells, platelets and other types of leukocytes. 2. c-kit enrichment As we want to work only with HSCs and progenitors, we have to get rid of all the other cells that were in the bone marrow we sampled. That's why we do an enrichment: this doesn’t purify as well as FACS sorting would, but you get way more HSCs and progenitors than other types of cells. To do this, we do a MACS (Magnetic-Activated Cell Sorting) enrichment for CD117 (c-kit). C-kit is a receptor tyrosine kinase protein that is expressed on HSCs and progenitors. To enrich the CD117 positive cells (expressing c-kit), we add superparamagnetic (a form of magnetism in nanoparticles [1]) beads coupled to anti-CD117 antibodies which will bind to CD117. As the beads are on c-kit positive cells, those cells will get stuck in the ferromagnetic spheres of the column, while all the others will pass through the column. Once the cell suspension has flown through we wash 3 times to be sure we wash away as many CD117 negative cells as possible. Finally, the column can be removed from the magnet: as the magnetic field isn’t retaining the beads anymore, we can get our CD117+ cells by flushing the column. Here you can see what this enrichment does: Before enrichment we had only 2% c-kit positive cells. CD117 enrichment almost multiplied it by 7 and gave us 13.7% c-kit positive cells. So here you see that there still are a lot of c-kit negative cells in the sample, the enrichment doesn’t purify it, it just amplifies the cells we want. 3. Antibody staining Once the c-kit positive cells are selected, we can go into deeper identification by adding other antibodies. To do this, we prepare a cocktail of different antibodies that will bind differentially on multipotent progenitors (MPPs) and HSCs. As I told you before, c-kit is expressed on all HSCs and progenitors. Sca-1 (stem-cell antigen 1) is another protein expressed on stem and progenitor cells [2]. Using flow cytometry, we can select the cells that are both positive for c-kit and sca-1 (LSK cells). So you can see here that c-kit enrichment doesn’t give a pure population: a lot of cells express neither sca-1 nor c-kit, meaning they’re not HSCs and progenitors. From this LSK population, we can then identify which cells are MPPs and HSCs. For this we use anti-Flt3R and anti-SLAM antibodies. As I told Cécile in my last post, Flt3R is a cytokine receptor that is expressed only on MPPs surface in the LSK gate. Inversely, SLAM are markers identifying HSCs [3]. That’s how we can find out what cells are MPPs and HSCs: Flt3R+ SLAM- are MPPs, Flt3R- SLAM+ are HSCs. Once you have the gates, you can tell the FACS to sort only cells from these gates. After about an hour of sorting, it will get you one tube of pure HSCs and another of MPPs ! References
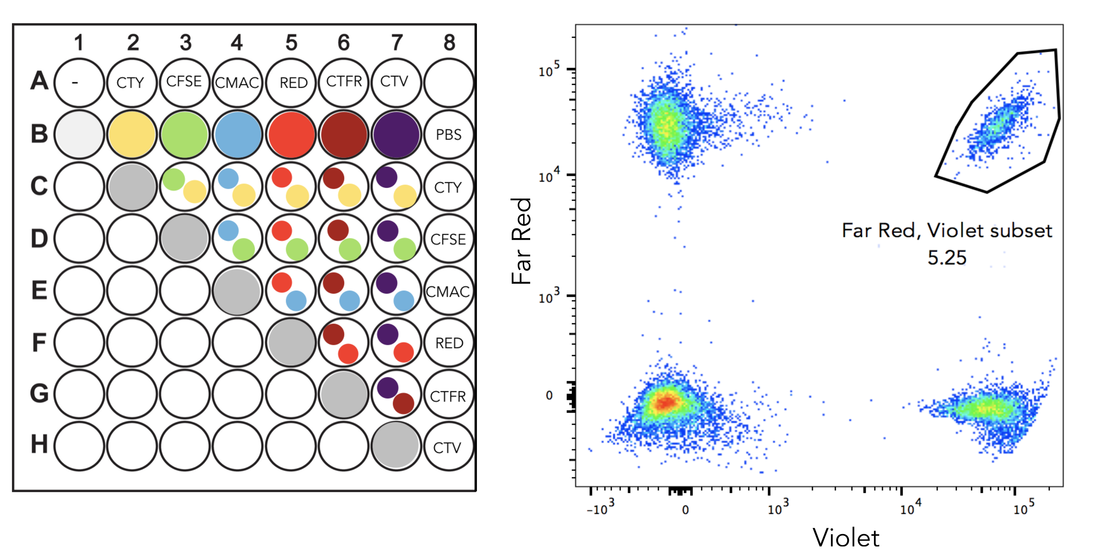
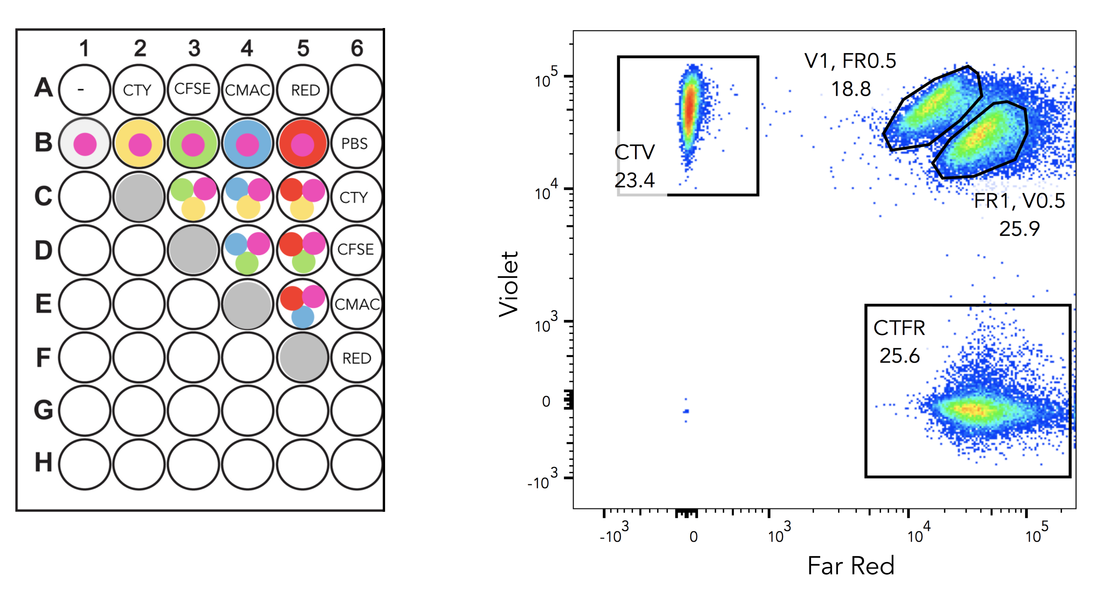
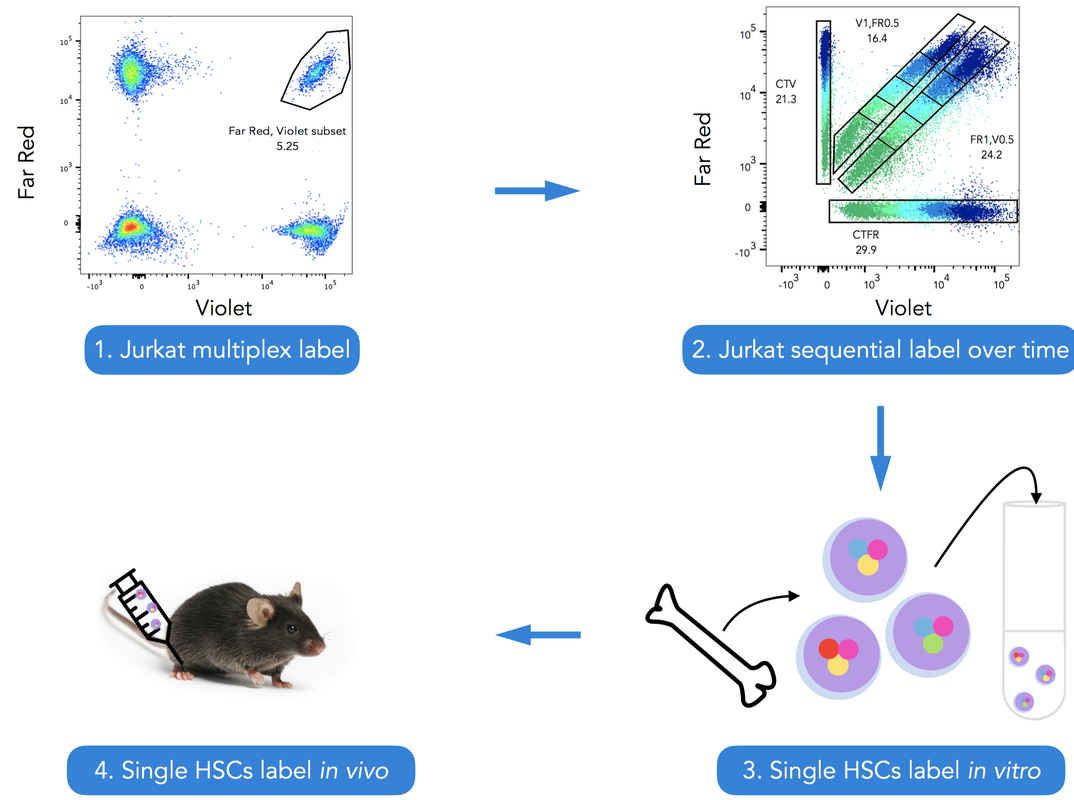
[1] https://en.wikipedia.org/wiki/Superparamagnetism [2] Holmes, Christina, and William L. Stanford. "Concise Review: Stem Cell Antigen-1: Expression, Function, And Enigma". Stem Cells 25.6 (2007): 1339-1347 [3] Yilmaz, O. H. "SLAM Family Markers Are Conserved Among Hematopoietic Stem Cells From Old And Reconstituted Mice And Markedly Increase Their Purity". Blood 107.3 (2005): 924-930 Credits Black6 mouse: https://www.jax.org/strain/003548 Bone and syringe: https://thenounproject.com Column: www.miltenyibiotec.com Now that you know what my team does, I will present you my project here. Question The aim of the project is to track proliferation of single stem cells in vivo, using fluorescent proliferation dyes. The two questions leading the experiment are: 1. What is the level of concordance of stem cell division ? (i.e. do both daughter cells divide the same way, synchronous, and does each daughter cell divide as often?) 2. How many times does each injected stem cell divide ? Hematopoietic stem cells (HSCs) differentiation has already been studied at a population scale with fluorescent dyes, but following single cells is new. Fluorescent cell staining dyes Before going into the details of my project, I’ll give you some important information about the staining dyes. The most well-known of them is CFSE (carboxyfluorescein succinimidyl ester) and it’s often used in immunology. It binds covalently to intracellular amine-containing molecules[1], resulting in homogeneously labeled cells. When a labeled cell divides, it gives 1/2 of its fluorescence to each daughter cell. Then they give 1/4th of the fluorescence to their daughter cells and so on. Theoretically, these dyes allow detection of up to 8 cell divisions before the fluorescence is too low to discern from background (when the dye is too diluted to detect its fluorescence). It’s a great tool to follow cell proliferation: as the cells have different amounts of fluorescence given their number of division, it’s possible to observe how many times they have divided (Figure 1). 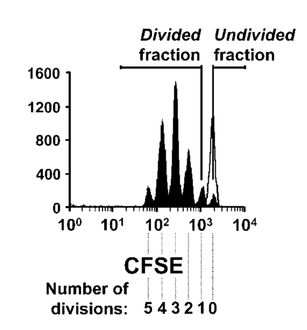 Fig.1: Fluorescence of the cells after 2 days culture Fig.1: Fluorescence of the cells after 2 days culture Here's an example of an experiment with CFSE labeled cells. Each peak represents a population of cells. The more they divide, the more the fluorescence decreases, that’s why the undivided fraction (population of undivided cells) peak is the highest. Method 1- Multiplex label on Jurkat cells Before working with HSCs, it’s easier to use a cell line. A cell line offers 2 main advantages: it’s easy to use because its cells are immortal and proliferate constantly. What’s more, we don’t have to kill a mouse each time we need cells. That’s why we developed the staining method using Jurkat cells, a human T lymphocyte line, stemming from a young boy who had T cell leukaemia. To follow single stained stem cells, each cell has to be labeled differently. We have 6 different dyes: CTY (CellTrace Yellow), CFSE, CMAC (CellTracker Blue CMAC), CellTracker Red CMPTX, CTFR (CellTrace Far Red) and CTV (CellTrace Violet). However, 6 different colors is not enough: if we inject more than 6 stem cells in a mouse, we can get more information by sacrificing only one animal. That’s why we first decided to combine the 6 colors 2 by 2 to get more unique stains. For now on, I will call this “multiplex label”[2] and here is what my 96 well plate looked like: The population on the top right is positive for Far Red and Violet. With this method, we can get 22 different combinations out of only 6 stains, which is a good start but not enough yet. 2- Sequential label on Jurkat cells, over time So we thought about some more combinations. To get them, a second layer of label is added to the multiplex label, the “sequential label”[3]. So we selected 2 colors, Celltrace Violet (CTV) and Celltrace Far Red (CTFR), and labeled the cells in 4 different concentrations: CTV 1:2000 CTFR 1:2000 CTV 1:2000, CTFR 1:4000 CTV 1:4000, CTFR 1:2000 After this first labelling, I quenched (stopped) the staining and pooled everything: here there are 4 different types of labeled cells in only one tube. Then, I can use these cells (represented by pink dots on the figure) to make a multiplex label: As you can see of this figure, there is a new population. Both cell subsets on the top right are double positive for Far Red and Violet, but at different concentrations. As the sequential label gives 4 different combinations and the second label 11, their combination gives us 4x11= 44 different combinations with only one mouse, while we would need 44 mice if we had one color !! Once the labelling technique was working well, I started following the labeled Jurkat cells over time, to see if I could see some proliferation going and for how long we could detect fluorescence. 3- Single HSCs label in vitro This is the stage I am right now. Here I’m doing the same thing I did with Jurkat cells, but on HSCs taken from murine bone marrow. Then, they must be activated to proliferate and grow in medium in vitro. After a couple of days, I will analyse the sample using flow cytometry and see if the cells have proliferated and how many times each cell divided. 4- Single HSCs label in vitro The last part of the project will be to do the same in vivo. First, take stem cells from a donor mouse, label them and inject the 44 differently labeled cells in another mouse. Before the injection, the mouse will be sub lethally (twice less than lethal) irradiated to kill half of its immune system (so that our HSCs can proliferate). After a few days the mouse is sacrificed, the bone we injected the cells in is sampled and analysed to find back our cells and see how they divided. If you’re a bit confused, here’s a short recap of my project: References:
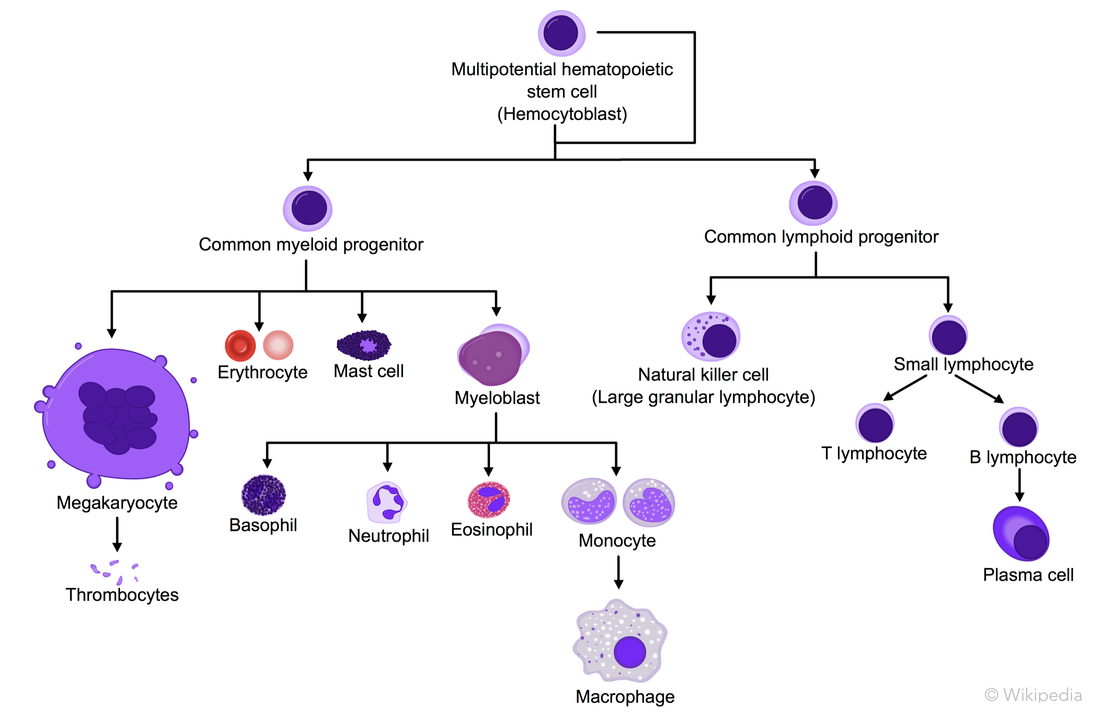
[1] Parish CR (December 1999). "Fluorescent dyes for lymphocyte migration and proliferation studies". Immunology and Cell Biology. 77 (6): 499–508. doi:10.1046/j.1440-1711.1999.00877.x. PMID 10571670 [2] Andersen, Rikke Sick et al. "Parallel Detection Of Antigen-Specific T Cell Responses By Combinatorial Encoding Of MHC Multimers". Nature Protocols 7.5 (2012): 891-902. [3] Marchingo, J. M. et al. "T-Cell Stimuli Independently Sum To Regulate An Inherited Clonal Division Fate". Nature Communications 7 (2016): 13540. Credits: Fig.1: Oostendorp, Robert A. J., Julie Audet, and Connie J. Eaves. "High-resolution tracking of cell division suggests similar cell cycle kinetics of hematopoietic stem cells stimulated in vitro and in vivo." Blood 95.3 (2000): 855-862. Black6 mouse: https://www.jax.org/strain/003548 Bone and syringe: https://thenounproject.com Everything else: my skills Where ? Quantitative immuno-hematology team, Curie Institute, Paris What’s the aim ? To understand at a single cell level how multipotent hematopoietic stem cells (HSCs) differentiate into white blood cells, involved in immune response, during hematopoiesis. Hematopoiesis is the process by which blood cells derive from a hematopoietic stem cell. Here is the classical hematopoiesis model, with a stem cell that can either self-renew, not to run out of HSCs, or differentiate to other cell types. From there on, there are several differentiation steps until mature white blood cells are produced. During these steps the HSCs lose their potency: once a cell has differentiated into a common lymphoid progenitor, it cannot become a macrophage anymore. Traditionally, the hematopoietic tree is represented like this: However, numerous studies have shown this model might not be correct in vivo[1]. For example, some cell types are missing on this figure and some cell subsets were thought to be closely related, turned out not to be. So the aim of my team is to understand this hematopoietic tree in vivo, that is to say, to clarify each cell’s lineage. How ? By using single cell tracing methods, as well as bioinformatics and mathematical modelings. Why should we care ? Because diseases affecting blood cells often lead to an excess or a lack of one cell type. As immune response is a team effort of many different white blood cells, missing one can have a dramatic health impact. Thus, unraveling the lineage of each cell type is essential to understand those diseases and acquire a more precise knowledge about our immune system. What about my project ? I will use fluorescent cell stains and flow cytometry to follow single cells over time and understand how HSCs differentiate to become blood cells. First, I’ll do this in vitro and then in vivo. I’ll give you more details about this in further posts. [1] Rathinam, Chozhavendan, and Richard A. Flavell. “The Hematopoiesis Paradigm: Clarity or Ambiguity?” Blood 112.9 (2008): 3534–3535. PMC. Web. 2 Feb. 2017. |