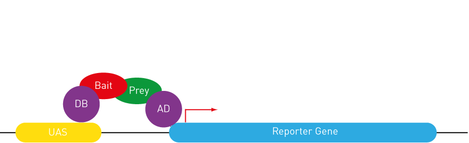
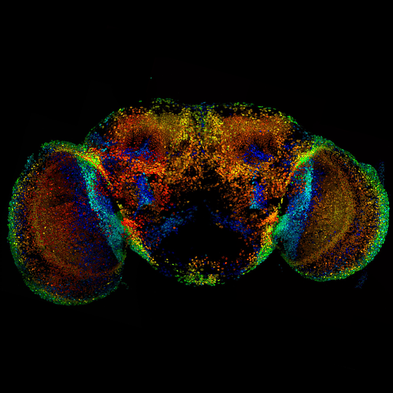
understanding the function of lola for newly formed neurons maintenance: how flies can help (Cécile)18/3/2017 Lola is a transcription factor required to maintain newly formed neurons in a differentiated state and prevent them from reverting to the neural stem cell type initiating tumors. This function implies the repression of neural stem cell genes and the activation of other genes’ transcription. So, Lola must interact with other factors to ensure opposite functions. Identification of potential co-factor for Lola The yeast two hybrid assay is very useful to determine protein-protein or protein-DNA interactions, by tracking the resulting downstream effects. The reporter gene chosen must encode a measurable transcription product, which is produced if a functional GAL4 binds to the Upstream Activation System (UAS). The principle of the assay is to split GAL4 in two fragments and attach a protein of interest to each. The first one contains the binding domain (BD) recognizing the UAS sequence. The second one contains the activating domain (AD) initiating transcription. Transcription can only be induced if the both fragments are recruited. In Lola’s case, Lola was used as a Bait, attached to GAL4 BD. A large number proteins from a drosophila DNA library was tested as Prey, attached to GAL4 AD. When the transcription product was observed, the protein from the library was considered as a candidate co-factor for Lola. Lola can interact with a protein recruiting the transcription machinery Rcd5 was detected through the yeast two hybrid assay. This protein is involved in the regulation of gene transcription by facilitating the recruitment of RNA Polymerase II Complex to the promoter region of target genes (1). Understanding how Lola fulfill its function could therefore benefit from investigating its interaction with Rcd5. As any previous research has been done, I will first practice a staining of Rcd5 using fluorescent antibodies within adult Wild Type (1118) Drosophila brains. Although the distribution of this essential protein is expected to be ubiquitous over the brain, I will maybe observe up-regulation in specific cell type… Following this general overview, I will use a Drosophila line over-expressing Rcd5 to get more information about protein function, although the precise experiments has not been designed yet. A protein involved in degradation of RNA can interact with Lola, to modulate its function? The yeast two hybrid assay has revealed many other proteins likely to interact with Lola. As I explained it to you on my previous blog post, removal of Lola only causes dedifferentiation of immature neurons. It means that more permanent mechanisms are recruited to repress neural stem cells genes throughout development. So, some factor might bind to Lola to modulate its function. I will be investigating one of them, the Zcchc7 protein, which appeared to be involved in the nuclear exosome. This complex acts as a quality-control system that degrades non-functional RNA but also regulate normal mRNAs and repress viral RNA in the nucleus (2). Nothing is known about its interaction with Lola and Zcchc7 could both enhance and repress expression. I began to investigate the role of this interaction using a drosophila line in which the sequence encoding this protein were down-regulated by RNAi in the Nervous System only. However, staining adult brains with antibodies did not reveal any tumours or abnormal brain development. This might be due to the low efficiency and lack of specificity of RNAi. In any case, antibodies staining gives amazing pictures, and I wanted to show you one from my lab, there are few other on the website… Crossing flies to direct the expression of target genes
I am also setting up a crossing. In Females genotype, GAL4 protein encoding sequence is placed under the control of promoters naturally expressed in neurons and neuroblast only. In Male genotype, a small hairpin RNA targeting Zcchc7 encoding sequence has been inserted. As expression of this interfering RNA requires GAL4 to bind the UAS, Zcchc7 downregulation will only occur in neurons and neuroblast in the progeny for which I will conduct a survival assay throughout embryos development. I will maybe try to stain some adult brains to compare this way of silencing gene expression with my previous results. Insert genes of interest into flies: construct the plasmids first Finally, I am constructing 3 plasmids that will be infiltrated into flies. The first contains a GFP tag to localize the Zcchc7 encoding sequence within the tissue. The second one is a control without GFP to ensure that the tag does not affect drosophila physiology. The last one contains a fusion protein: once flies are transfected, the sequence encoding Zcchc7 protein also encodes Dam methylase. This enzyme methylate at low level the molecular targets of the protein! The technique is called TaDa and has been set up by the lab I am working on itself (3). However, I have to deal with several issues while trying to make the plasmids, mainly due to the large size of the insert (5 kB). References
6 Comments
Noémie Paillon
22/3/2017 04:23:15 am
Reply
Cécile
22/3/2017 09:39:06 am
It was quite random : they used a cDNA library of Drosophila Melanogaster Larvae brain, 56.7 millions interactions were tested, 165 colonies, 111 with a good confidence score and those with the highest score are being investigated now.
Reply
Noémie Paillon
23/3/2017 03:07:31 am
That must have been quite a lot of pipetting !!!!
Reply
Cécile Crapar
23/3/2017 05:27:28 pm
Yes I totally agree for non coding RNA that is why I am talking about aberrant RNA, could be both noncoding and coding
Reply
margaux
31/3/2017 02:58:27 am
Hi Cecilus,
Reply
Cécile Crapart
1/4/2017 08:53:55 am
Hi! The advantage of using confocal is that you can observe slice of an entire brain. I have to observe the brain I stained at 60 magnification to reach the cell level, I'll tell you more once it will be done!
Reply
Leave a Reply. |