 Fig.1: let's start with a bit of humour, shall we? Fig.1: let's start with a bit of humour, shall we? Relax and enjoy your flight on Biostatistics Airlines: First thing I’ve done while crossing path with medical biostatistics was… laughing. A mirthless laugh: those formulae were unintelligible. What I did not understand then was that, at our little level, the only thing we need is… knowing what we want! Indeed, each described situation can be reduced to a formula. In the previous post, we’ve seen that distributing patients into prognostic strata was possible by comparing two genes mutation status within different tumour types and their combined impact on prognostic. What is the prognostic here? Simply survival. What do we want? Simply knowing if Mr X with mutated A but wild-type (i.e the gene “normal” state) B will live longer than Mrs Y with both wild-type A and B. So let’s not forget that statistics can be easy to grasp if those concepts are explained with the right handle [1] and let’s slowly take one step after another to understand how survival analysis is made and can help us on our journey toward personalised medicine. First step: datum datum datum, dati, dato dato, DATA, DATA, DATA, datorum, datis datis We are all familiar with the first step of every statistical analysis: data collection. In the introduction, it is said that we are interested by survival: at a given time is a given patient dead or alive (yes, mathematics can be pretty cynical)? Some of you who already know a bit about statistics may ask why we do not use the classical models such as the Normal Law (Gaussian curve [2]) that allow easy interpretations. It is because survival do not follow classical distributions and events are often distributed as follows: several early events, some late events [3]. So survival analysis requires specific models. There are two “events” that can be observed: death or recurrence (i.e cancer comes back). Models use two variables linked to these events: i) overall survival: time between diagnostic and death or the last follow-up ii) progression free survival: time between the response to the treatment and the recurrence or the last follow-up Here “follow-up” means the time when you examine all the patients in your cohort. What is interesting is that this notion of follow-up is another point that makes survival analysis different from other models. If patients do not exhibit death nor recurrence, they are considered as “censored” in the study i.e they do not give objective data therefore classical models are not relevant either. The only thing you need to keep in mind is that censoring events follows specific rules [4]. Second step: “All Curves Are Beautiful” (Yvan Cornut) Now that we have our data, we have to put them into functions to model what we want. Two functions are used [3]: i) survival S(t) that represents the “cumulative non occurrence” i.e the the summed time a patient survived ii) hazard h(t) that represent the event occurrence at a given time t knowing that until t, the patient did not experience any events. Those function will be useful to plot survival curves, a way to represent survival in order to analyse it more easily. The most common way to do it is by using the Kaplan-Meier method: 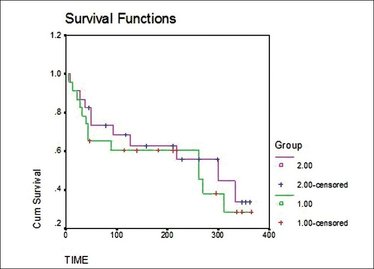 Fig.3: Kaplan-Meier curves, S(t) = f(t) Fig.3: Kaplan-Meier curves, S(t) = f(t) As initial conditions we have:
If you’re a cynical, you can plot [1-S(t)] vs. time to obtain death curves but let’s stay positive, after all, you’re still alive after the previous explanations :)
Next step: comparing your curves. Why? Because all differences are not significant. Huu... what? Let me be clearer: when you cook, it will be significantly different to add 10 more grams of butter than only one gram. Same thing for your survival curve: How big must be the difference so you can conclude that if Mr X with wt-B and mutated A live longer, everyone is more likely to also live longer? To do that, we use a test named log rank [6] and performed the same way as a χ² [5]. No, no, don’t go! I’ll be happy to develop in the comments for those who may be interested but let’s stop there on survival curves and take another step. Last-but-not-least step: what else? In the first blogpost, we saw that cancer was a multifactorial disorder with internal and external causing factors. Methods like the ones above (Kaplan Meier and log rank) are univariate. Hence the need for models that take into account the factors linked to the patient (remember: personalised approach) named cofounders or covariates and that allow to estimate clinically [7] (not statistically) the impact of the mutation status combined with those factors on the prognostic. There are two categories of models that differ by the way impact on the prognosis is conceived. In the Cox proportional hazard model that we will see below, each factor has a weight that influences the prognosis whereas in the Accelerated Failure Time model that will not further be detailed, each factor can shrink or stretches the survival time along the time-axis. Briefly, the Cox semi-parametric is a way to link event occurrences with the covariate set. It uses the following function of the hazard h(t): h(t) = h0(t) x exp(i=1pbi.xi) That can appear absolutely barbaric until you just know that p is the number of covariates you are considering (age at diagnosis, gender, other disorder…), x a given covariates and b its relative coefficient that weigh the hazard. Still following? Great! Here are some little precisions: i) h0(t) is a “basal hazard” that is very convenient because we do not have to assume that h(t) follows a given and known distribution ii) This model can only be applied under the assumption that all the risks are constant multiples. iii) exp(bi) is the risk ratio and allows to better understand the model: if b>0, h(t) increases therefore, if the risk increases, the survival time decreases. Thus we have a negative correlation between risk ratios and survival Now you know! Congratulations, you have succeeded in following statistical explanations! What to remember and tell your family during dinner so they consider you as some kind of wizard? First, survival analysis are made following three steps: data collection, survival curves plot and comparison, estimation of covariates impact on the prognostic groups defined by your curves. Second, what matters in biostatistics is to know what you want because only then will you know what hypotheses to make and what test or model to use to answer your question. Here are presented tests and models that are “non parametric” which mean that you assume your data do not fit a given distribution. This is convenient because survival distributions are often different from what exists but parametric test are considered more robust therefore, looking at what can be done with parametric models is interesting too. Finally, I really hope that you were able to follow everything above because that would mean that I was able to simplify those notions and that you maybe understand the most important message to take home: statistics can be easily understood and even if we won’t be biostatistician, it is important to understand what we read in papers. The only way to be legitimate in our criticism is by knowing what we’re talking about :) Références: [1] Aberkane, I. (2016). Libérez votre cerveau !. 1st ed. Paris: Robert Laffont. [2] Mathsisfun.com. (2017). Normal Distribution. [online] Available at: https://www.mathsisfun.com/data/standard-normal-distribution.html [Accessed 24 Feb. 2017]. [3] Clark, T., Bradburn, M., Love, S. and Altman, D. (2003). Survival Analysis Part I: Basic concepts and first analyses. British Journal of Cancer, 89(2), pp.232-238. [4] Bradburn, M., Clark, T., Love, S. and Altman, D. (2003). Survival Analysis Part III: Multivariate data analysis – choosing a model and assessing its adequacy and fit. British Journal of Cancer, 89(4), pp.605-611. [5] Chi Square Statistics. (2017). [online] Math.hws.edu. Available at: http://math.hws.edu/javamath/ryan/ChiSquare.html [Accessed 22 Feb. 2017]. [6] http://www.oxfordjournals.org/our_journals/tropej/online/ma_chap12.pdf [7] Bradburn, M., Clark, T., Love, S. and Altman, D. (2003). Survival Analysis Part II: Multivariate data analysis – an introduction to concepts and methods. British Journal of Cancer, 89(3), pp.431-436. Figures:
6 Comments
Hortense
1/3/2017 07:26:11 am
Dear Margaux ;)
Reply
Margaux
2/3/2017 07:09:29 am
Dear Hortense,
Reply
margaux
2/3/2017 07:15:50 am
I just found out how to explain the tricky thing about the two hypotheses
Aurélien DIEHL
1/3/2017 01:13:19 pm
Hi Margaux!
Reply
margaux
2/3/2017 08:02:03 am
I was going to answer your first question but in my sources, you have this great website http://math.hws.edu/javamath/ryan/ChiSquare.html that takes a huge advantage over my answer: you’ll find tables that are far more explanatory than any words I could try to write about it.
Reply
margaux
2/3/2017 09:00:27 am
Ho I should have been more explicit: i is just the index of your covariate. For example, let’s say that you have three covariates: Leave a Reply. |

