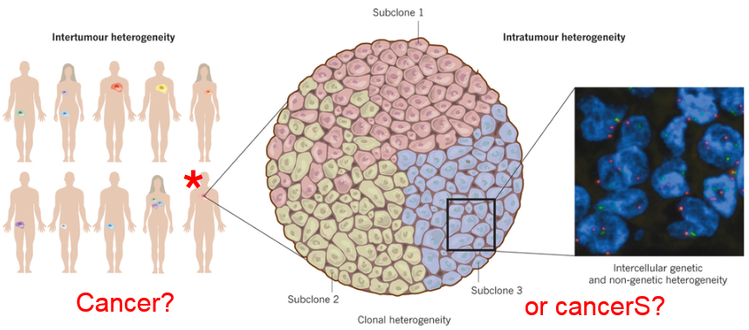
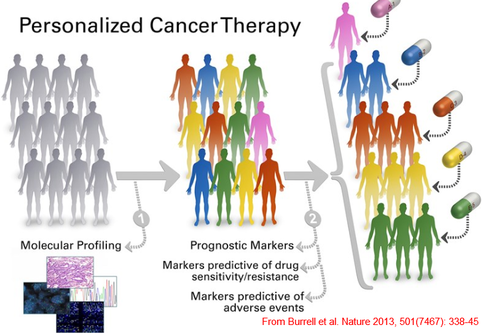
 Why is personalised medicine a different approach? Personalised medicine is a paradigm shift: from the quest of a treatment that will fit most of the patients, the aim is now to tailor treatments to the patient’s medical history, personal “parameters” (age, gender...), environment, even microbiome (one’s microbial population might be as unique as one’s fingerprints [1]). Most of the diseases are diagnosed by looking at the phenotype but, sometimes, two sets of symptoms can overlap and characterising a disorder becomes complex. A more personalised approach can be useful as factors responsible for the phenotype (i.e symptoms) can differ and looking at individual factors could lead to easier characterisation thus better treatment of the disorder. Personalising is supposed to maximise the benefits and minimise side effects [2] but it is an ideal to pursue and in practice, available technologies and knowledges do not allow to fully achieve it. Nonetheless, research to personalize medicine offers fascinating examples and, in the oncology field in particular, allows new tumour classifications, first step towards a complete personalised characterisation. Here is outlined what can be done in oncology thanks to these new technologies. One important part of the personalised medicine is the genomic medicine [3] i.e the medicine that tries to extract information from genomes (structural genomics) or from transcriptomes/proteomes/epigenomes (functional genomics). Such approach is now possible because of the Next Generation Sequencing that allows a higher throughput (i.e the number of nucleobases read by run by day)and oncology benefited from this huge leap forward. Characterising tumours is like taking measurements for a custom-made tuxedo: Cancers used to be characterized by an organ-based classification [2] but genetic analysis allowed to discover that cancers resulted from an accumulation of mutations and some of those mutations were found within many neoplastic tissues: an organ-based classification became obsolete. Moreover, tumours are composed of several sub-clonal populations and healthy cells that interact so there is intra-heterogeneity (differences within the same tumour). Finally, two patients with the same tumour location can have different subclones so there is inter-heterogeneity (differences between two individual tumours from the same type) too. Alexandrov et al. [4] studied 4,938,362 mutations from 7, 042 cancers and that ideally allows to look for two things:
Describing is a start… But predicting matters too: There was an important word in the previous sentence. Until now, we only saw how genetic analyses allow to characterise a disorder: this is the diagnosis. BUT! One aim of a personalised treatment is also to PREDICT. How can we do that in practice is one of the subjects of my internship on gliomas and can be briefly explained step-by-step: i) two types of tumours are studied to study the prognostic variation; ii) two genes are studied with one that is known to be frequently found in those tumours or known to play a role in the tumorigenesis and the second one that will help to categorise; iii) correlations between clinical and genetic observations: patients are separated into several groups according to their mutation status and clinical data are coupled to each group. For example, if patients with genes A and B mutated live longer than those with A mutated and normal B, clinicians will be able to predict the disease evolution in other patients with known mutation status. Another interest: if a specific mutation-status-associated phenotype is known to be susceptible to react to a drug, a mutated patient can be treated with it (crucial when we know chemotherapeutic side effects). Now you know! What is the take-home message? First, new sequencing technologies allow to study diseases from another point of view and to open new therapeutic windows. Second, the more precisely a disease is characterised, the more tailored the treatment can be and this is why personalised medicine development is tightly correlated to genetic advances. Third, personalised medicine is in its infancy: a completely tailored treatment is rare and one limitation is that personalised treatments must also take into account the resistance that can arise (for example, radiotherapy in cancer selects the strongest phenotypes and in many cases of recurrence, the tumour is less likely to be destroyed) and the evolution of the disease so medicine will truly be personalised when we will understand better how disorders work. Finally, even if it is the beginning, personalised medicine allows to improve diagnosis/prognosis in several fields including cancer by taking the first step before a complete individual characterisation: the stratification of the patients into risk/prognostic groups. REFERENCES, in case you’d like to learn more: [1] Franzosa, E., Huang, K., Meadow, J., Gevers, D., Lemon, K., Bohannan, B. and Huttenhower, C. (2015). “Identifying personal microbiomes using metagenomic codes”. Proceedings of the National Academy of Sciences, 112(22), pp.E2930-E2938. [2] Ogino, Shuji et al. “Cancer Immunology—Analysis Of Host And Tumor Factors For Personalized Medicine”. Nature Reviews Clinical Oncology 8.12 (2011): 711-719. Web. 8 Jan. 2017. [3] Ginsburg, Geoffrey S. and Huntington F. Willard. "Genomic And Personalized Medicine: Foundations And Applications". Translational Research 154.6 (2009): 277-287. Web. 8 Jan. 2017. [4] Alexandrov, Ludmil B. et al. "Signatures Of Mutational Processes In Human Cancer". Nature 500.7463 (2013): 415-421. Web. 8 Jan. 2017. [5] Boisselier, B., Gallego Perez-Larraya, J., Rossetto, M., Labussiere, M., Ciccarino, P., Marie, Y., Delattre, J. and Sanson, M. (2012). “Detection of IDH1 mutation in the plasma of patients with glioma”. Neurology, 79(16), pp.1693-1698. [6] Hanahan, Douglas and Robert A. Weinberg. "Hallmarks Of Cancer: The Next Generation". Cell 144.5 (2011): 646-674. Web.
4 Comments
Amélie
20/2/2017 07:26:30 am
Hello Margaux,
Reply
Margaux
21/2/2017 01:45:41 am
Hi Amélie!
Reply
Noémie
21/2/2017 04:16:25 am
Hi Margaux !
Margaux
21/2/2017 04:28:11 am
Yes indeed, you're perfectly right!
Reply
Leave a Reply. |