|
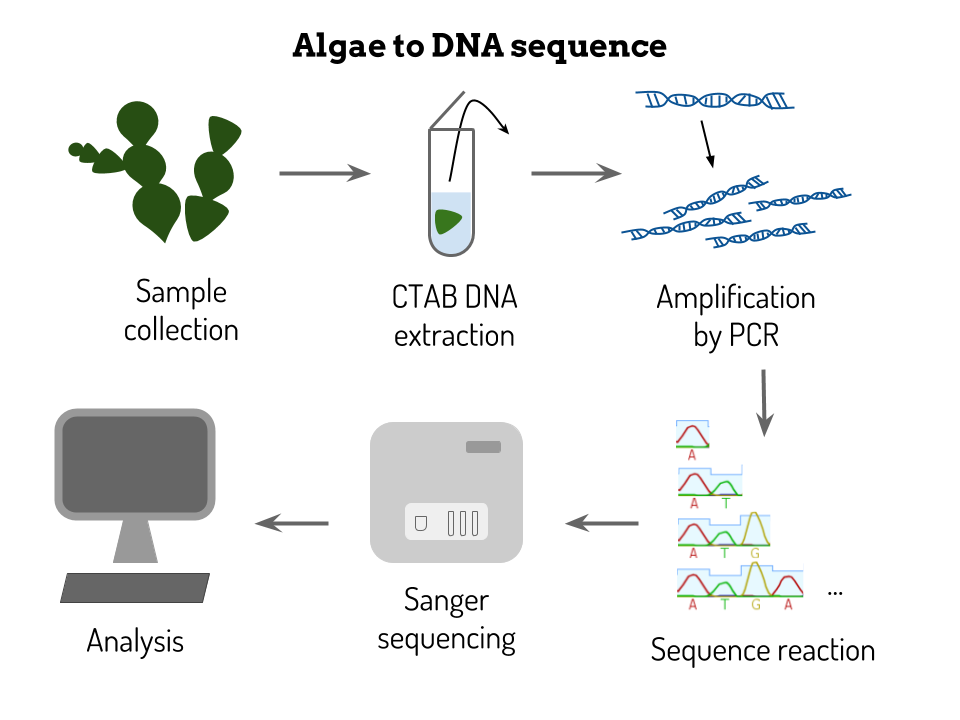
My internship has been composed of two main parts : a laboratory part and a bioinformatic analysis part! During my lab work, I started with a small marine macroalgae sample and obtained in the end a DNA sequence. The techniques I used are widely used for molecular analysis in biotechnology: DNA extraction, DNA amplification by PCR and finally DNA sequencing by the Sanger Method. DNA extraction DNA is extracted from the algae cell's nuclei, there is just one multi-nucleic cell, by using physical force as well as chemical compounds to degrade the cell wall. I used a ball mill to physically tear up the algae cell's tissue. By putting a little stainless steel ball inside the microtube with my tissue sample and then shaking the tube at a high frequency, what is left in the tube no longer looks like a little algae, but is rather a greyish green powder. We then add a lysis buffer composed of CTAB and Proteinase K to chemically degrade the tissue. A four hour lysis, during which the sample is heated at 60°C and agitated at 800 rpm, also uses a physical force, heat, to degrade the cell walls. Finally, this solution is purified thanks to a mix of chloroform and isoamyl alcohol which separates the DNA from the protein and lipid compounds. DNA amplification We then dose the DNA solution using a NanoDrop, corresponding to a small spectrometer, to know how much DNA was extracted from the cell. The DNA we obtained is then amplified by Polymerase Chain Reaction (PCR) on certain regions of interest, the tufA and rbcL markers. For a quick recap about PCR you can check out this very clear video. We therefore used existing primers to amplify these sequences of interest and visualized results by electrophoresis. If the results were not satisfactory, meaning that we had a double mark or no mark at all, we modified the DNA solution's concentration or the PCR program. Sanger Sequencing Once we have obtained a highly concentrated DNA solution with only the fragment we are interested in sequencing, we can purify the solution using a Sephadex column. The solution is then ready to be sequenced, starting with the sequence reaction using only one primer. Sanger sequencing is resumed here. Since sequencing is automated by a machine, the only part where I manipulated was during the sequencing reaction. I have therefore obtained a chromatogram with my sequence, now on to the bioinformatics part ! On my computer, more magic happens! Sequence analysis Once I have obtained my chromatogram, I correct certain ambiguities thanks to the Geneious program. Like Margaux had shown in her previous post, sometimes nucleotide peaks can be overlapping and in that case, the analyst has to decide whether one is more prominent or whether it remains inconclusive. The sequences also need to be trimmed (most commonly at the beginning and at the end) to keep only our region of interest as well as only the legible part of the sequence. Once the sequences are clean, I can align them all using the muscle alignment method. I then use this alignment to create several different kinds of phylogenetic trees following the Maximum Likelihood reconstruction method as well as the Neighbor Joining method. I compare these results and analyze two things : what species clade my samples correspond to and if they are genetically close to other samples from the Indian Ocean or not. This helps me see both the number of different species that were found during the sample collection as well as validate or invalidate existing patterns and hypotheses of speciation. Here is a little recap in visual form : You'll hear more about my results soon !
Don't hesitate to ask any questions you might have!
6 Comments
Margaux
20/4/2017 04:10:43 am
hello Clara!
Reply
Clara
22/4/2017 04:29:51 am
Thank you for your questions Margaux!
Reply
Margaux
25/4/2017 01:03:32 am
Perfectly well! Thanks :)
Cécile
24/4/2017 03:13:55 am
Hi!
Reply
Clara
25/4/2017 04:38:46 pm
Hi Cécile!
Reply
Cécile
28/4/2017 07:22:48 am
Thanks I understand well now! Leave a Reply. |