|
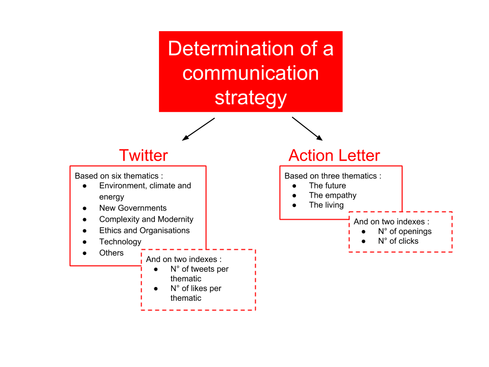
At Engage, I am responsible for communication. The aim of this mission is to continuously update the network about Engage’s activity and develop a communication strategy in order to satisfy audience’s interests. For this reason, I developed a study for determining the thematics that interest the most Engage’s audience. To do so, I took into consideration the number of “likes” and posts I shared on each thematic on Twitter and on the thematics of the three newsletters (called Action Letters) I wrote based on the numbers of openings and clicks on it (Fig. 1). The results will determine the subjects of the following events organised by Engage. Figure 1: The thematics and the indexes taken into consideration on Twitter and on the Action Letter for the determination of the subjects of the next events organised by Engage.
0 Comments
Some details of the process by which a hematopoietic stem cell becomes a mature immune cell remain unclear to this day. To tackle this issue, a new approach that already brought lots of new information is used: studying hematopoiesis at a single-cell level. It is to that extent that the project of my internship aims at developing a new tool to study single-cell dynamics in hematopoiesis. To do this, we design a new cell staining method using multiple combinations of proliferation dyes. It allows to follow the differentiation of 44 single hematopoietic stem cells as well in vitro as in vivo.
Today, I will talk to you about a problem we are currently facing without having the solution yet. When water is taken from the sea the amount of incoming salt – Total Dissolved Solids (TDS) - will be in a short range of values allowing us to determine easily what should be the solution used not only for pre treatment but also for the kind of semi permeable membrane to be used. However, this is not the case of the plant I am working on. We have a TDS range between 500 mg/L (maximum value of World Health Organisation for drinking water) and 42 000 mg/L (sea water). Therefore, quoting smartly and to minimizing the power consumption is a challenge.
An article published in desalination explains the parameters to accommodate such variability are the membrane selection, the vessel configuration, the energy recovery device, dual pumping system and extra product water storage capacities. This post will explain you why each of the parameters is a way to deal with such TDS and how it works. 1. Membrane selection In one vessel, there are usually 7 membranes, all of the same kind. Or, if we use less membrane in one vessel and a two pass system –meaning the brine is going to be treated also and the permeate of the brine will be mixed with the permeate of the first round – as well as different kinds of membranes in one vessel, this will distribute the productivity through the vessel allowing better recovery. 2. Vessel configuration As said in the previous paragraph, 7 is usually the number of membranes in a vessel. However, it is not mandatory and can be changed. Therefore, if we decreased the number of membrane it will decrease the outlet feed but it will also decrease the TDS in the brine allowing for more stages and better recovery. 3. Energy recovery device The energy recovery device will allow less consumption of energy as it will reduce the number of High Pressure Pumps, which are the most energy demanding equipment. 4. Dual pumping system The dual pumping system is about adding a pump with a lower pressure output so that if the salt amount in the water is less, the water will by-pass the high pressure pump and go to the low pressure pump. This will allow energy to be saved. 5. Extra product storage capacity The extra product storage will allow diminishing the salinity of the water by blending the stored water to the outlet if the outlet water quality is not sufficient. However, it mainly works for regular tide cycle. I hope this was clear to you. As my internship is coming to end, it is the best time to present you in a hundred words what I did during almost four months.
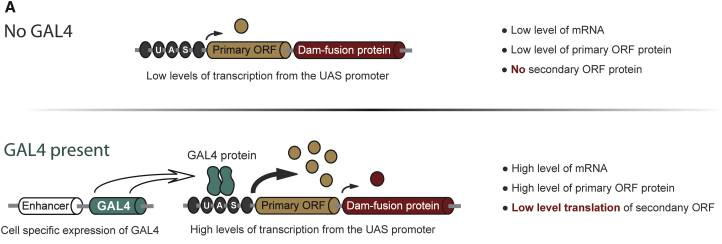
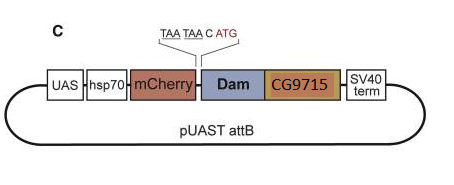
My internship was about ALS. I worked on cellular functional project and mainly on a genetic bio-informatics project. The first one was about the effect of CSF of ALS patients on motoneurons derived from IPSc. Unfortunately, I dropped the plate on the analysis day, preventing us to get results… One the second hand, I did a genetic association cases-controls study. The goal was to associate new genetic variants to ALS. To do so I used a cohort of 46 ALS patients from Europe that I compared to a cohort of controls from Europe. I use a collapsing statistic test named Optimal Sequence Kernel Association Test (SKAT-O). The test worked but unfortunately it did show any significant results. I learned a lot in this internship and it gave me a willing to work in bio-informatics. Aurélien I am doing my internship in a lab which has developed a technique for identifying the transcription factors expressed in a cell-specific manner. In other words, it gives information about the genes being transcribed in the cell-type you are studying, such as neurons or neuroblasts, at different stage of development, such as embryonic or adult stages. This is very interesting because as I explained in my first blog post, stem cells differentiate to specific fate following a transcriptional program of the genome. Unlike other existing methods, this technique called TaDa can be used in vivo and does not require cell purification. Methylation as a tag to identify the genetic targets of a protein More generally speaking, the method allows us to identify the genes which are bound by a protein of interest for any model organism used (and the while genome is screened). This is achieved through the fusion of an enzyme, DNA adenine-methyltransferase from Escherichia coli, to the protein of interest. The genes with which the protein of interest interacts will be identified and tagged by a methyl group, attached by the enzyme, to the adenine base of the DNA. The fusion protein is placed under the control of UAS activation system so that tissue-specific expression can be achieved using the GAL4 promoter (GAL4 is placed under the control of promoter naturally expressed in neurons, neuroblasts or other specific cells). You can find more information about the GAL4/UAS system in my previous blog post. Methylation: a toxic tag? However, Dam methylase must be expressed at very low level because high methylation of DNA can be toxic for the organism or nonspecific. The issue at this point was thus to reduce levels of Dam methylase translation and avoid toxicity. Because ribosomes can reinitiate transcription, the lab split the construct with a codon stop. This codon is located after the UAS and the primary Opening Reading Frames to reinitiate and reduce levels of translation of the Dam-fusion protein (see below on fig.2). Hence, on the one hand, the primary ORF (ORF1 in the drawing) is translated at high levels. This is interesting as it encodes a fluorophore, mCherry, allowing us to locate the construct within cells. On the other hand, the secondary ORF encoding Dam methylase is supposed to be translated at low levels after reinitialization of translation (Fig.1). Without GAL4 being expressed, the primary ORF only would be translated at low level. So, the genetic targets of the protein of interest will be tagged in a cell-type specific manner. The lab demonstrated that this construct was not toxic at any stage of development. Is the technique accurate? They have assessed the accuracy of their new technique using RNA Polymerase II as protein of interest as it has been studied many time in previous studies. TaDa technique accurately identified RNA Pol II occupancy in the genome when compared with other methods! I will use this technique! One of the goal of my project is to use this technique to determine the genetic targets of the Zcchc7 protein, involved in the nuclear exosome and interacting with Lola. I am thus constructing a plasmid where Dam is fused to Zcchc7 so that the DNA bound by Zcchc7 will be tagged by methylation (Fig.2)! Reference
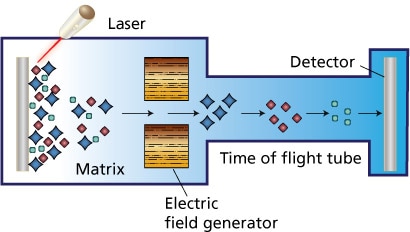
Southall, T.D., Gold, K.S., Egger, B., Davidson, C.M., et al. (2013) Cell-type-specific profiling of gene expression and chromatin binding without cell isolation: Assaying RNA pol II occupancy in neural stem cells. Developmental Cell. [Online] 26 (1), 101–112. Available from: doi:10.1016/j.devcel.2013.05.020. So, as you might remember, my job during my internship is taking care of the Accreditation of the newly arrived mass spectrometer. But what is a mass spectrometer? A mass spectrometer is a device that detects the protein composition of a component. In microbiology, it detects the ribosomal proteins contained in a certain micro-organism and, comparing the profile obtained to a data bank, identifies the micro-organism. There are different types of mass spectrometry and they differ from one another by their way of tracking and their way of detecting the analyzed sample. The mass spectrometer I worked on is a MALDI-TOF type of mass spectrometer. MALDI-TOF stands for Matrix Assisted Laser Desorption/Ionization - Time of Flight. A sample is mixed with a charged matrix (either with protons or electrons; in my particular case, charged with protons) that crystallizes on the sample and inserted in the spectrometer. A laser hits the sample, causing the ribosomal proteins to get charged/ionized by the protons of the matrix and get air-born. They then pass by an electromagnetic Field that accelerates them. The particles are then sent through a tube towards a detector. Thanks to the time the particles take to arrive to the detector, their weight can be deduced. Here is what a specter looks like: Thanks to the specter, you can deduce what proteins are present and in what proportions. How did the mass spectrometer come to be? Well, both K.Tanaka et al (Japan) and Karas & Hillenkamp (Germany) are credited for its invention, though only K.Tanaka got a Nobel Prize for it (2002). We could also go further backwards and credit blablabla……. This method of identification is fairly new in microbiology. How where micro-organisms identified before? The answer is observing Gram staining, observing the appearance of the colonies and using biochemical and agglutination tests. Agglutination tests rely on ….. and biochemical tests allow to identify the behavior of a certain micro-organism when in the presence of this biochemical ….. The mass spectrometer may seem like a failsafe method, but it has its faults. Here are a few of them: - A specter may not be characteristic of the sample analyzed. By this, I mean that it may correspond to multiple samples and, as such, may cause difficulties in analysis and force to use an alternative method to differentiate. - You have to be sure of the purity of your sample References:
This blog post will be less scientific because I wanted to talk about the startup and how we work. In fact, it is quite different than in research laboratories. In this post, I will also talk a bit about our teamwork, and something important to me, which is “Do what you like to do”. TeamworkAs a startup, we have a team of designers, who imagine and create decorations and scenography. There is also the team for laboratory research, who modifies genetically the strains, tries different methods of production… And finally, there are the heads, who make decisions for the startup and for example decisions for special events. Every week we have a team meeting, where we talk about future events, one of us presents his last results, someone else presents a scientific article which can be interesting for Glowee, and often someone brings some “pains au chocolat”, essential for a good brain activity. ;) Moreover, we often share our ideas or problems about laboratory work or others things, thus we can find solutions or give hints to other team members. Of course, there are stressing moments, when we prepare an event, or just because we are a startup and we need to have good results (to be able to continue this adventure), but we work in a very good atmosphere, and everyone is part of the adventure! No typical days in GloweeFor the designers as well as for the lab team, every day is different. I will talk more about the lab because this is where I work ;). As I told you last time, I work on cloning and bioluminescence production in bioreactors. In a week I can work on both, or only one, it depends on the results, on the different needs for the events (sometimes we have to try some things before an event), but it also depends on the ideas of my supervisor and me. In fact, taking our results into account, we think about the different tests, for the bioreactor with 2 different culture media for example. I also spend time on Arduino (of course you know what it is but for those who'd need a little reminder, you'll find it here https://www.arduino.cc/ ), in order to make a light reference, with a LED and an Arduino UNO, for the experiments. We needed a light reference to analyse the pictures taken during the bioluminescence production. With this, we can see the accuracy of the camera. So, finally, every day are different from each other! Do what you like to doIt is true that “Do what you like to do” is more easy to say than to realize in certain situations or in certain countries. But I think this is essential to you, to your work, to your life and how you feel every day. I mean, wake up the morning and say “let’s go, it will be a good day” is sooo important. It is true that in certain families or in certain countries it is more difficult to access to the university you want or you're dreaming job. Unfortunately, it can be limited by money or just the diversity of schools and companies. But honestly, for those living in occidental countries we have a lot of opportunities and possibilities. So, when you found the courses, the internship, the job or whatever you like, just GO! For those who have less opportunities, if you are motivated, a lot of things are possible, never give up. Also, it is never too late to change your job or learn something new! To conclude, do what you like to do guys, it is so important. Why I did this post Still, in internship at Glowee, I really appreciate Glowee’s work, the team, what and how I learn new things. So, I wanted to share my experience and my vision of the startup work with you. Hope you completely like what you do!
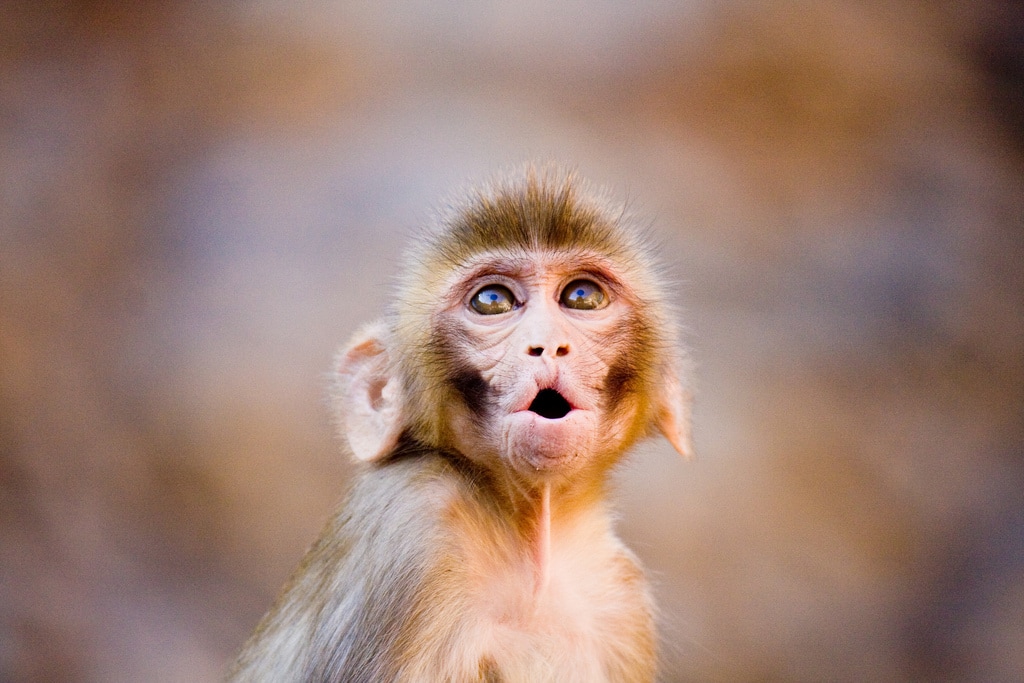
In my laboratory, each Tuesday we have a Lab Meeting of one hours with all people . This week, we spoke of Animals Ethics ! For remember, i realize an internship at Stem cell and Brain Research Institute. Like many laboratory we have animals for the research. So this Lab Meeting was very particular about this sensible subject. In the room of conference there was searchers on stem cell, brain, robotics, administrative people and veterinary, most people had divergence points of view on the subject. This day, the director of research of my lab wanted to speak of new proposition* concerning the large public. Today, we can look horror pictures on animals laboratory on youtube, facebook or other web site. So we formed in our mind a picture of this subject very delicate, a horror vision ! I don’t say this vision is not True, but I want today share with you an other version more adapted for the animals ethics. In Neuroscience we study several subject on animals, each new research protocol which include laboratory animals is presented front of ethic committee which give a validation or not to start the project, in fonction of several criteria ! The criterias can change in function of each project, but keep the same main way. In neuroscience each variation of emotion, or changement of physical aspect on the subjects can have negative effect on the experiment. So if you want a good results, we have to pay attention for our animals. In my team, each animals have a name, games, times and a specially relation with her searcher and a veterinary. So, why inflict a bad treatment on the laboratory animals, when we can have more without that ? * I can't speak of this proposition now ! Fig 1 - Rhesus Monkey
Bibliography :
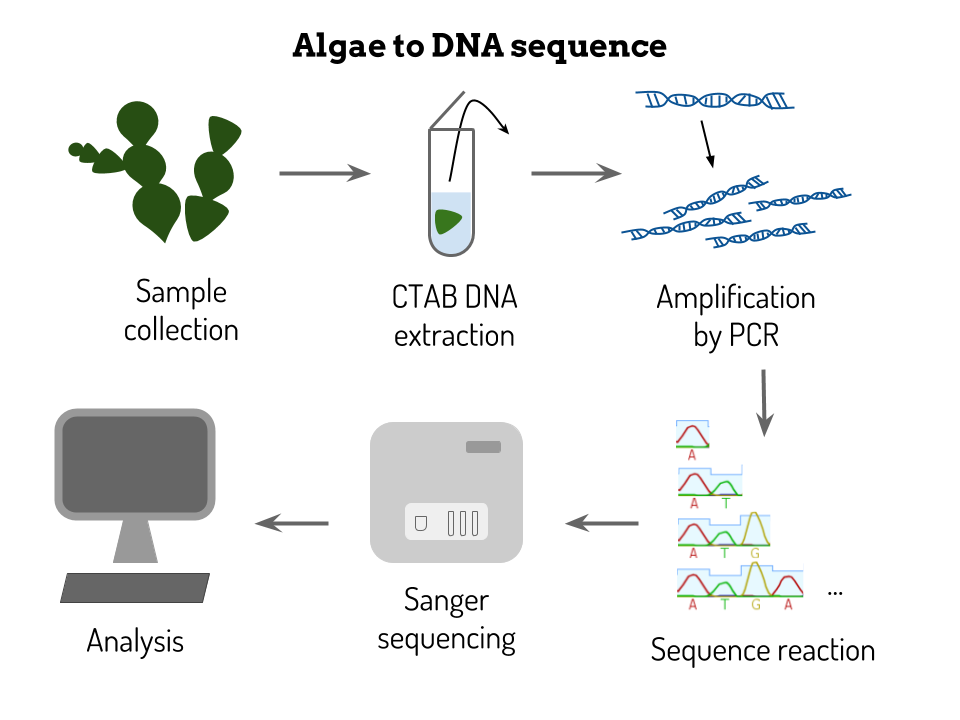
My internship has been composed of two main parts : a laboratory part and a bioinformatic analysis part! During my lab work, I started with a small marine macroalgae sample and obtained in the end a DNA sequence. The techniques I used are widely used for molecular analysis in biotechnology: DNA extraction, DNA amplification by PCR and finally DNA sequencing by the Sanger Method. DNA extraction DNA is extracted from the algae cell's nuclei, there is just one multi-nucleic cell, by using physical force as well as chemical compounds to degrade the cell wall. I used a ball mill to physically tear up the algae cell's tissue. By putting a little stainless steel ball inside the microtube with my tissue sample and then shaking the tube at a high frequency, what is left in the tube no longer looks like a little algae, but is rather a greyish green powder. We then add a lysis buffer composed of CTAB and Proteinase K to chemically degrade the tissue. A four hour lysis, during which the sample is heated at 60°C and agitated at 800 rpm, also uses a physical force, heat, to degrade the cell walls. Finally, this solution is purified thanks to a mix of chloroform and isoamyl alcohol which separates the DNA from the protein and lipid compounds. DNA amplification We then dose the DNA solution using a NanoDrop, corresponding to a small spectrometer, to know how much DNA was extracted from the cell. The DNA we obtained is then amplified by Polymerase Chain Reaction (PCR) on certain regions of interest, the tufA and rbcL markers. For a quick recap about PCR you can check out this very clear video. We therefore used existing primers to amplify these sequences of interest and visualized results by electrophoresis. If the results were not satisfactory, meaning that we had a double mark or no mark at all, we modified the DNA solution's concentration or the PCR program. Sanger Sequencing Once we have obtained a highly concentrated DNA solution with only the fragment we are interested in sequencing, we can purify the solution using a Sephadex column. The solution is then ready to be sequenced, starting with the sequence reaction using only one primer. Sanger sequencing is resumed here. Since sequencing is automated by a machine, the only part where I manipulated was during the sequencing reaction. I have therefore obtained a chromatogram with my sequence, now on to the bioinformatics part ! On my computer, more magic happens! Sequence analysis Once I have obtained my chromatogram, I correct certain ambiguities thanks to the Geneious program. Like Margaux had shown in her previous post, sometimes nucleotide peaks can be overlapping and in that case, the analyst has to decide whether one is more prominent or whether it remains inconclusive. The sequences also need to be trimmed (most commonly at the beginning and at the end) to keep only our region of interest as well as only the legible part of the sequence. Once the sequences are clean, I can align them all using the muscle alignment method. I then use this alignment to create several different kinds of phylogenetic trees following the Maximum Likelihood reconstruction method as well as the Neighbor Joining method. I compare these results and analyze two things : what species clade my samples correspond to and if they are genetically close to other samples from the Indian Ocean or not. This helps me see both the number of different species that were found during the sample collection as well as validate or invalidate existing patterns and hypotheses of speciation. Here is a little recap in visual form : You'll hear more about my results soon !
Don't hesitate to ask any questions you might have! When I did research about bioinformatic I found a blog that was very helpful and taught me a lot. When I wanted to know more about this blog, I found an incredible story about its author Eric Vallabh Minikel and his wife Sonia The story begins when Sonia’s mother died at the end of 2010. The biopsy of Sonia’s mother revealed a mutation in the PRNP gene on the chromosome 20. This mutation is known to be associated to the disease that caused the death of Sonia’s mother, the prion disease. Prion disease is also named Creutzfeldt-Jakob disease, Fatal Familial Insomnia (FFI) or the mad cow disease. It is a dominant mutation that causes the prion disease around the age of 50. Dominant means that Sonias had 50% chances to carry it too. On December 2011, Eric and Sonia finally receive the diagnosis, the test was positive which means that Sonia carries the same mutation as her mother. At this moment, their lives changed, the quest of their life to cure Sonia just begun. Sonia, who was only 27, was graduated from Harvard Law School and had a job as consultant and Eric was working as a software engineer and data analyst in transportation sector. They created the Prion Alliance an association for the research into the treatment or cure of prion disease. They began to take night classes, go to conference, and read papers about the disease and finally left their old career to find job in Massachusetts General Hospital’s Center for Human Genetic Research, Eric as an informatician and Sonia as a stem cell technician. At the same time, Eric started the biology blog http://www.cureffi.org After receiving a price for their research on prion disease in 2014, the couple were admitted to a PhD program in biological and biomedical sciences in the Harvard Medical School. In May 2015, they were invited to joined the lab in the Broad Institute of MIT and Harvard. From the start of their quest, this awesome couple still did not find a cure to the prion disease but they already found clues and initiated the Prions@Broad project. In 2016, they also were present for the Obama’s signature of 21st Century Cures Act, a “legislation that provides for investments in research and development, and delivery of treatments for disease.” Beyond this beautiful love story of this couple that left their old life to start a quest to cure Sonia, we can see an atypical motivation for doing biomedical research. Sonia and Eric are in this field for something else than academic results. Sometimes during my internships, I felt like there was a gap between research field and the clinical reality that leads to an abstraction of the disease. I saw that sometime the seek for research is to get a great academic recognition through publications. Should not it be for something else? For the good of research maybe we should reconsider our motivation? Through his blog, Eric shares his experience, contribute to improving his field and helps one those who encounter the same difficulties. Aurélien
|